Volume 15, Number 5—May 2009
Research
Virulent Strain of Hepatitis E Virus Genotype 3, Japan
Cite This Article
Citation for Media
Abstract
Hepatitis E virus (HEV) genotype 3, which usually causes asymptomatic infection in Japan, induced severe hepatitis in 8 patients. To better understand genetic features of HEV associated with increased virulence, we determined the complete or near-complete nucleotide sequences of HEV from these 8 patients and from 5 swine infected with genotype 3 strain swJ19. Phylogenetic analysis showed that the isolates from the 8 patients and the 5 swine grouped separately from the other genotype 3 isolates to create a unique cluster, designated JIO. The human JIO-related viruses encoded 18 amino acids different from those of the other HEV genotype 3 strains. One substitution common to almost all human HEV strains in the JIO cluster was located in the helicase domain (V239A) and may be associated with increased virulence. A zoonotic origin of JIO-related viruses is suspected because the isolates from the 5 swine also possessed the signature V239A substitution in helicase.
Hepatitis E virus (HEV) infection is relatively common. Anti-HEV antibodies are found in 10%–20% of the general population in Japan and most Asian countries (1,2); however, only a small fraction of these infections induce overt hepatitis. Although the mechanisms underlying induction of liver damage by HEV have not been well characterized, HEV genotypes seem to have distinct disease-inducing potential. HEV sequences have been classified into 4 genotypes (3). Genotype 1 consists of epidemic strains in developing countries of Asia and Africa. Genotype 2 is represented by the prototype sequences from an epidemic in Mexico, which have also recently been detected in Africa. Genotypes 3 and 4 are distributed worldwide and have been implicated in sporadic cases of acute hepatitis E in humans and domestic pigs. HEV genotypes 3 and 4 are found in Japan, but fulminant or severe acute hepatitis develops more frequently in persons infected with genotype 4 (4–6). The severity of liver disease may therefore be influenced by the HEV genotype with which the patient is infected as well as host factors such as age, gender, and pregnancy status.
In 1997, we identified a strain of HEV from a patient in Japan who had acute hepatitis (designated JIO) that clustered with genotype 3 sequences. From 2004 through 2006, JIO-related viruses were isolated from 7 additional patients who had acute or severe hepatitis. To better understand genetic features of HEV associated with severe hepatitis, we compared the complete or near-complete sequence of JIO isolates from these 8 patients with other well-characterized genotype 3 and 4 isolates. To determine whether these human genotype 3 sequences were zoonotic in origin, we sequenced full-length viral genomes from 5 swine infected with the swJ19 strain of HEV. These 5 animals were part of a larger outbreak of HEV infection that occurred in swine at a single farm in southern Japan during 2000–2002. The GenBank/EMBL/DDBJ accession numbers for nucleotide sequences of HEV isolates are AB291951–7/AB291960 (for the human isolates) and AB443623-7 (for the swine isolates).
We enrolled 8 patients who were infected with HEV genotype 3 and had clinical features of hepatitis (Table 1). A zoonotic source of HEV infection was identified for 3 of these patients: pig liver for patient 4, pig meat for patient 6, and wild boar meat for patient 7. Prothrombin time, a surrogate marker of hepatic insufficiency, averaged 63.9% (± 29.1%) of the reference range among the 8 HEV genotype 3–infected patients. Hepatitis was particularly severe in patients 3, 5, 7, and 8; at the peak of disease, prothrombin times for these patients ranged from 27% to 46% of the reference range. These sporadic HEV cases were not clustered geographically; they were distributed across several regions of Japan, including southern (Okinawa) and northern (Saitama) prefectures (Figure 1). Informed consent was obtained from all patients after the nature and purpose of the study was explained to them.
To assess possible zoonotic origins of these human infections, we sequenced HEV strain swJ19 isolates from 5 of 11 swine with previously documented infections (7). These animals had been raised commercially at a farm in the southern part of Miyazaki Prefecture where HEV infections were detected during 2000–2002. All animals received humane care, and the study was approved by the institutional review committee of Toshiba General Hospital, Tokyo, Japan.
To determine whether infections could be linked to a common genotype 3 virus, we compared the genetic structure and sequence homology of 8 human and 5 swine HEV strains. The entire or near-complete nucleotide sequences of the 8 JIO strain isolates from the human patients and the swJ19 strain isolates from the 5 swine were determined by a method reported previously (8,9), with some modifications. In brief, nucleic acids were extracted from serum with the QIAamp MinElute Virus Spin Kit (QIAGEN GmbH, Hilden, Germany). HEV RNA genomes were reverse transcribed, and cDNA was amplified by PCR with primers specific for 23 overlapping regions of the HEV genome. Reverse transcription and first-round PCR were conducted by using the SuperScript III One-Step RT-PCR System (Invitrogen Corporation, Carlsbad, CA, USA); second-round PCR was conducted with the Platinum Taq DNA polymerase (Invitrogen). The 5′- and 3′-terminal sequences were amplified by using the SMART RACE cDNA Amplification Kit (Clontech Laboratories Inc., Mountain View, CA, USA) and Oligo (dt)20 Primer (Invitrogen), respectively. The sequences enriched in G-C were amplified with the TaKaRa LA Taq in GC Buffer (TaKaRa Shuzo Co. Ltd., Shiga, Japan). The sequences not amplifiable by the above PCR methods were subjected to PCR with primers deduced from adjacent 5′ and 3′ sequences. The final products were sequenced in the 377 DNA Sequencer with use of the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA, USA). Genetic analyses of HEV sequences were conducted by the unweighted pair-grouping method with arithmetic means by using computer software GENTYX-MAC Version 13 (Genetyx Corporation, Tokyo, Japan).
The prototypical isolate, JIO-Sai97L, had a genome length of 7,215 nt that contained a 5′ untranslated region (UTR), 3 open reading frames (ORFs), a 3′ UTR, and a poly-A tail. Lengths of HEV genomes from 6 other patients (JYM-Tot04L, JYU-Oki04L, JSS-Oka04L, JIY-Tot05L, JSO-Oki05L and JSW-Kyo-FH05L) were identical to that of JIO-Sai97L. An exception was the HEV isolate JTK-Kag06C from patient 7, which was slightly longer (7,236 nt). The 5 HEV isolates from swine (swJ19-1, swJ19-2, swJ19-5, swJ19-7, and swJ19-8) had genomes of 7,210 nt. The 3 ORFs of all swine and human HEV genomes had identical protein coding capacity. HEV isolates from all human patients had 97.9%–98.6% sequence homology with the prototypical JIO-Sai97L strain from patient 1. The 5 swine swJ19 isolates had 98.3%–99.9% sequence homology when compared with each other and 98.0%–99.8% homology when compared with the JIO strain from human patients.
Comparison of nucleotide sequences of the 13 human and swine HEV isolates in this study with those of published HEV genotype 3 sequences showed that the 13 complete and near-complete sequences described in this study closely matched those of 2 well-characterized genotype 3 viruses: JRA1 (89.4%–89.7% nucleotide identity) and swJ570 (88.9%–89.0% nucleotide identity). The 13 human and swine genotype 3 isolates displayed weak homology with other HEV genotypes. The B1 isolate of genotype 1 (GenBank accession no. M73218) was only 74.1%–74.7% similar to these genotype 3 viruses, the M1 isolate of genotype 2 (accession no. M74506) was only 73.6%–74.0% similar, and the T1 isolate of genotype 4 (accession no. AJ272108) was only 75.6%–76.0% similar.
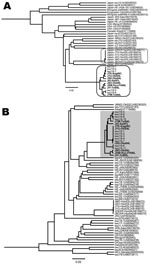
Using the 13 complete or near-complete genomic sequences of HEV genotype 3 isolates described in this study (Figure 2), we constructed a phylogenetic tree. HEV sequences from the 8 patients (JTK-Kag06C, JYU-Oki04L, JSS-Oka04L, JIO-Sai97L, JSO-Oki05L, JSW-Kyo-FH06L, JIY-Tot05L, JYM-Tot04L) clustered on a branch separate from the other genotype 3 sequences, forming a distinct grouping related to the prototypical JIO strain. The swJ19 HEV sequences from the 5 swine (swJ19-1, swJ19-2, swJ19-7, swJ19-5, and swJ19-8) clustered closely with the JIO-related viruses from the human patients, indicating that the human and swine HEV isolates were highly similar (Figure 2, panel A). Another 18 swine isolates, from farms other than the 1 involved in the swJ19 outbreak, were phylogenetically distinct from those of the outbreak farms (Figure 2, panel B).
Another genotype 3 cluster was formed by 6 isolates from Hyogo Prefecture in western Japan (Figure 2, panel A). In this cluster were 5 HEV isolates from persons in whom hepatitis developed after they ate uncooked deer meat (10) and from serum from a local boar and a deer (11). Unlike the JIO-related viruses, which were broadly distributed from the most southern to northern Japanese prefectures, HEV strains responsible for the infections in Hyogo Prefecture were not commonly found in other parts of the country. Broad distribution of the JIO-related viruses seems to be unique in HEV epidemiology. In 2 (25%) of these 8 patients, pig liver or meat had been implicated in HEV infection.
Comparison of the 13 JIO-related viruses (Figure 2, panel A) with the other genotype 3 strains also showed 18 aa differences: 12 in ORF1, 3 in ORF2, and 3 in ORF3 (Table 2). Three mutations in the JIO strain were characteristic of genotype 4 viruses, which are typically more pathogenic than other HEV genotypes. ORF1 differences were found at amino acids 605 (serine to proline, S605P), 978 (isoleucine to valine, I978V), and 1213 (valine to alanine, V1213A). The V1213A substitution is potentially most relevant because it was not found in the prototypical isolate from patient 1 (JIO), who had mild clinical disease when infected in 1997, but was present in highly related isolates from the other 7 patients who had more severe hepatitis during 2004–2006. V1213A in ORF1 corresponds to V239A of the helicase domain, and its surrounding sequences were well conserved in HEV isolates of genotypes 3 and 4 (Appendix Figure). Because V239A is common in genotype 4 isolates, we analyzed genomes of the genotype 3 JIO-related viruses for evidence of intergenotypic recombination. Comparison of 28 genotype 4 sequences with those of the JIO-related isolates showed no obvious signs of recombination (data not shown), which suggests that the V293A substitution arose independently in this genetically unique cluster of genotype 3 viruses. Notably, all 5 isolates recovered from swine on the Miyazaki Prefecture farm during the outbreak of 2000–2002 possessed the V239A substitution.
Circumstantial evidence indicates that HEV genotype influences the severity of liver disease. Remarkably, HEV seroprevalence studies in Egypt found no clinical illness in any person, including pregnant women, although most (67.7%–84.3%) had been exposed to HEV genotype 3 (13,14). In contrast, results of a survey of 254 patients with HEV infection in Japan showed hepatitis associated with genotype 4 to be more severe than that associated with genotype 3 (4). Our results showed a clustering of 8 HEV isolates of JIO strain, genotype 3, recovered from patients with clinical hepatitis.
Despite the high disease-inducing activity of the HEV JIO strain, the 8 patients infected with this strain were distributed widely over Japan. This distribution is at odds with a local cluster of genotype 3 infections restricted to persons with hepatitis and to wild animals living in Hyogo Prefecture, Japan (Figure 2, panel A) (11). Wide regional distribution has also been reported for some HEV genotype 4 isolates (15). Why JIO strains caused more severe hepatitis than might be expected for a genotype 3 virus is not clear, but the reason may depend on the magnitude of virus replication. Alternatively, recombination between divergent HEV strains (16) may have played a role. This possibility prompted us to look for any recombination of JIO strains with genotype 4 strains that cause severe hepatitis in Japan. However, we found no evidence of recombination between the JIO strain of genotype 3 HEV with which the 8 persons were infected and 28 isolates of genotype 4 retrieved from the public and our own databases. The 18 aa substitutions were unique to the 8 human JIO and 5 swine sw19 isolates and not present in other genotype 3 viruses. Three differences in ORF1 (S605P, I978V, and V1213A) were common in wild type genotype 4 but not in genotype 3 isolates (Table 2). Because S605P and I978V are located in an ORF1 region that has high sequence divergence, they are unlikely to be responsible for an enhanced disease-inducing capacity. In contrast, V1213A changes at amino acid 239 of helicase, an enzyme capable of enhancing the efficiency of viral replication (17), were detected in 7 of the 8 patients (Appendix Figure). Indeed, the helicase region of the prototypical JIO-Sai97L isolated in 1997 did not contain this amino acid polymorphism. Remarkably, all 5 swine isolates recovered in Miyazaki Prefecture during 2000–2002 belonged to the JIO strain and possessed V1213A (helV239A). Taken together, the evidence strongly suggests a zoonotic origin for the 8 human HEV infections with JIO-related viruses.
Experimental and circumstantial evidence suggests that helV239A may have enhanced the helicase activity of the genotype 3 JIO strain to levels comparable with those of the more pathogenic genotype 4 viruses. However, the role of helV239A in enhancing helicase activity should be evaluated in vitro in future studies; its role in inducing hepatitis is yet to be confirmed. In addition, the effect of other mutations of JIO strains need to be fully explored before a conclusion can be drawn regarding the hepatitis-inducing capacities of this strain of HEV.
Findings from this study have public health implications. Because farm swine constitute a melting pot for generating various HEV mutants, at least in Japan where virtually all swine become infected with HEV within 4 months of birth, it is conceivable that virulent HEV mutant(s) arise on pig farms. Such occurrence has been described for influenza, for which point mutations are associated with increased virulence (18,19); for example, mutant influenza viruses that arose on chicken farms in Hong Kong in 1997 were transmitted to humans and had fatal consequences (20,21). In addition, although a vaccine against HEV has recently been developed (22), a vaccination strategy for humans and animals has yet to be defined. The results of our study indicate that selective vaccination of farm swine bearing HEV isolates of high virulence, such as those of the JIO strain in Miyazaki Prefecture, should be recommended to decrease the incidence of fulminant or severe acute hepatitis E in Japan and elsewhere in the world.
Dr Takahashi is principal investigator in the Department of Medical Sciences at Toshiba General Hospital. His research interest is hepatitis viruses, most recently hepatitis E virus.
Acknowledgments
We thank Christopher M. Walker and Sheikh Mohammad Fazle Akbar for designing and checking the manuscript.
This work was supported in part by a grant from the Ministry of Health, Labour and Welfare of Japan.
References
- Arankalle VA, Tsarev SA, Chadha MS, Alling DW, Emerson SU, Banerjee K, Age-specific prevalence of antibodies to hepatitis A and E viruses in Pune, India, 1982 and 1992. J Infect Dis. 1995;171:447–50.PubMedGoogle Scholar
- Tanaka E, Matsumoto A, Takeda N, Li TC, Umemura T, Yoshizawa K, Age-specific antibody to hepatitis E virus has remained constant during the past 20 years in Japan. J Viral Hepat. 2005;12:439–42. DOIPubMedGoogle Scholar
- Emerson SU, Anderson D, Arankalle A, Meng XJ, Purdy M, Schlauder GG, Hepevirus. In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA, editors. Virus taxonomy: VIIIth report of the International Committee on Taxonomy of Viruses. London: Elsevier/Academic Press; 2004. p. 853–5.
- Abe T, Aikawa T, Akahane Y, Arai M, Asahina Y, Atarashi Y, Demographic, epidemiological, and virological characteristics of hepatitis E virus infections in Japan based on 254 human cases collected nationwide [in Japanese]. Kanzo. 2006;47:384–91. DOIGoogle Scholar
- Inoue J, Nishizawa T, Takahashi M, Aikawa T, Mizuo H, Suzuki K, Analysis of the full-length genome of genotype 4 hepatitis E virus isolates from patients with fulminant or acute self-limited hepatitis E. J Med Virol. 2006;78:476–84. DOIPubMedGoogle Scholar
- Ohnishi S, Kang JH, Maekubo H, Arakawa T, Karino Y, Toyota J, Comparison of clinical features of acute hepatitis caused by hepatitis E virus (HEV) genotypes 3 and 4 in Sapporo, Japan. Hepatol Res. 2006;36:301–7. DOIPubMedGoogle Scholar
- Takahashi M, Nishizawa T, Miyajima H, Gotanda Y, Iita T, Tsuda F, Swine hepatitis E virus strains in Japan form four phylogenetic clusters comparable with those of Japanese isolates of human hepatitis E virus. J Gen Virol. 2003;84:851–62. DOIPubMedGoogle Scholar
- Takahashi K, Iwata K, Watanabe N, Hatahara T, Ohta Y, Baba K, Full-genome nucleotide sequence of a hepatitis E virus strain that may be indigenous to Japan. Virology. 2001;287:9–12. DOIPubMedGoogle Scholar
- Takahashi K, Kang JH, Ohnishi S, Hino K, Miyakawa H, Miyakawa Y, Full-length sequences of six hepatitis E virus isolates of genotypes III and IV from patients with sporadic acute or fulminant hepatitis in Japan. Intervirology. 2003;46:308–18. DOIPubMedGoogle Scholar
- Tei S, Kitajima N, Takahashi K, Mishiro S. Zoonotic transmission of hepatitis E virus from deer to human beings. Lancet. 2003;362:371–3. DOIPubMedGoogle Scholar
- Takahashi K, Kitajima N, Abe N, Mishiro S. Complete or near-complete nucleotide sequences of hepatitis E virus genome recovered from a wild boar, a deer, and four patients who ate the deer. Virology. 2004;330:501–5. DOIPubMedGoogle Scholar
- Erker JC, Desai SM, Schlauder GG, Dawson GJ, Mushahwar IK. A hepatitis E virus variant from the United States: molecular characterization and transmission in cynomolgus macaques. J Gen Virol. 1999;80:681–90.PubMedGoogle Scholar
- Stoszek SK, Abdel-Hamid M, Saleh DA, El Kafrawy S, Narooz S, Hawash Y, High prevalence of hepatitis E antibodies in pregnant Egyptian women. Trans R Soc Trop Med Hyg. 2006;100:95–101. DOIPubMedGoogle Scholar
- Stoszek SK, Engle RE, Abdel-Hamid M, Mikhail N, Abdel-Aziz F, Medhat A, Hepatitis E antibody seroconversion without disease in highly endemic rural Egyptian communities. Trans R Soc Trop Med Hyg. 2006;100:89–94. DOIPubMedGoogle Scholar
- Takahashi K, Okada K, Kang JH, Karino Y, Ichida T, Matsuda H, A lineage of hepatitis E virus within genotype IV, associated with severe forms of hepatitis [in Japanese]. Kanzo. 2005;47:389–90 [cited 2009 Mar 6]. Available from http://www.jstage.jst.go.jp/article/kanzo/46/6/46_389/_article
- van Cuyck H, Fan J, Robertson DL, Roques P. Evidence of recombination between divergent hepatitis E viruses. J Virol. 2005;79:9306–14. DOIPubMedGoogle Scholar
- Ahola T, den Boon JA, Ahlquist P. Helicase and capping enzyme active site mutations in brome mosaic virus protein 1a cause defects in template recruitment, negative-strand RNA synthesis, and viral RNA capping. J Virol. 2000;74:8803–11. DOIPubMedGoogle Scholar
- Hatta M, Gao P, Halfmann P, Kawaoka Y. Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses. Science. 2001;293:1840–2. DOIPubMedGoogle Scholar
- Claas EC, Osterhaus AD, van Beek R, De Jong JC, Rimmelzwaan GF, Senne DA, Human influenza A H5N1 virus related to a highly pathogenic avian influenza virus. Lancet. 1998;351:472–7. DOIPubMedGoogle Scholar
- Subbarao K, Klimov A, Katz J, Regnery H, Lim W, Hall H, Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science. 1998;279:393–6. DOIPubMedGoogle Scholar
- Shrestha MP, Scott RM, Joshi DM, Mammen MP Jr, Thapa GB, Thapa N, Safety and efficacy of a recombinant hepatitis E vaccine. N Engl J Med. 2007;356:895–903. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 15, Number 5—May 2009
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Kazuaki Takahashi, Department of Medical Sciences, Toshiba General Hospital, 6-3-22 Higashi Oi, Shinagawa, Tokyo 140-8522, Japan
Top