Volume 16, Number 2—February 2010
Dispatch
Extensive Mammalian Ancestry of Pandemic (H1N1) 2009 Virus
Cite This Article
Citation for Media
Abstract
We demonstrate that the novel pandemic influenza (H1N1) viruses have human virus–like receptor specificity and can no longer replicate in aquatic waterfowl, their historic natural reservoir. The biological properties of these viruses are consistent with those of their phylogenetic progenitors, indicating longstanding adaptation to mammals.
In 2009, a new H1N1 influenza virus (pandemic [H1N1] 2009) emerged in Mexico, spread to the United States (1), and subsequently caused the first influenza pandemic of the 21st century (2). The emergence of pandemic (H1N1) 2009 virus is imperfectly understood, but an early switch in hemagglutinin (HA) receptor specificity is essential to allow interspecies transmission (3–5).
Pandemic (H1N1) 2009 virus strains were recently reported to be reassortants of the North American and European swine lineages (6). Phylogenetic evidence suggests that this reassortment event occurred 10–17 years ago (7). These data suggest that the current pandemic (H1N1) 2009 virus strains should have receptor specificity typically found in the HA of mammalian viruses (Neu5Acα2,6Gal). In addition, they may have lost the ability to replicate in avian hosts, the natural reservoir species. To test these hypotheses, we examined the biological properties of pandemic (H1N1) 2009 virus, including receptor specificity, erythrocyte binding, and ability to replicate in avian species.
We first tested species-specific erythrocyte agglutination by the pandemic (H1N1) 2009 isolates A/California/04/2009 and A/Tennessee/1-560/2009 and by other isolates from humans, swine, and birds (Table 1). The pandemic (H1N1) 2009 isolates showed reduced or absent agglutination of goose and chicken erythrocytes. Human and swine H1N1 viruses were agglutinated by turkey, guinea pig, chicken, and goose erythrocytes, and all erythrocytes we tested except those of swine were agglutinated by avian isolates (Table 1).
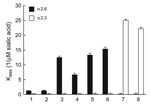
We next measured the receptor binding of the 2 pandemic (H1N1) 2009 isolates to sialic substrates, both natural (fetuin) and synthetic (3′-sialyllactose [3′SL] and 6′-sialyllactosamine [6′SLN] attached to a polyacrylic carrier) (Figure). The binding pattern to fetuin was identical among all isolates tested (association constant Kass ≈ 5.8 ± 0.5, 1/μM sialic acid). The currently circulating human and pandemic influenza (H1N1) viruses showed a preference for 6′SLN and negligible binding to the avian-type 3′SL. A similar pattern was observed for 2 recent swine viruses, which bound only to 6′SLN receptors with nearly equal affinity as pandemic (H1N1) 2009 isolates. As expected, the 2 avian H1 viruses bound strongly only to 3′SL (Figure).
To assess the infectivity and pathogenicity of pandemic (H1N1) 2009 virus strain A/California/04/2009 in terrestrial (chickens, quails) and aquatic (domestic and wild ducks) avian species, we inoculated 10 birds of each species by intranasal, intraocular, and intratracheal instillation with ≈106.0 of the 50% egg infectious dose (EID50) of the virus. We then observed the birds for the next 2 weeks for death and for viral shedding and signs of illness. No birds showed obvious clinical signs of disease. Virus was detected only on postinoculation day 1 in infected chickens and ducks and only in tracheal samples at low titers (<1.7 log10 of the EID50/mL [8]) (Table 2). However, no later shedding of virus was observed, indicating that the virus detected on postinoculation day 1 could have been caused by residual virus particles after inoculation. In contrast, our results revealed that the A/California/04/2009 strain efficiently infected quails with significantly higher titers (<3.4 log10 EID50/mL until postinoculation day 5; p<0.05) in both oropharyngeal and cloacal swab specimens (Table 2). The virus was detected in the trachea (1.7 log10 EID50/g), lungs (2.3 log10 EID50/g), and cecal tonsil (0.8 log10 EID50/g) of quails on postinoculation day 5.
The potential bird-to-bird intraspecies transmission of the A/California/04/2009 pandemic (H1N1) 2009 virus strain in avian species was also examined by introducing 3 contact birds to the inoculated birds’ cages on postinoculation day 1. There was no subsequent evidence of viral shedding through the upper respiratory tract or fecal-oral route in any group of birds except 1 of 3 contact quails (Table 2). Oropharyngeal virus titers in this quail were l.7 and 1.5 log10 EID50/mL on postinoculation days 3 and 5, indicating that productive viral replication was occurring.
The A/California/04/2009 pandemic (H1N1) 2009 virus strain showed minimal replication and no transmission in chickens and ducks (domestic and wild), but the virus replicated and had limited transmissibility in quails. Our finding is consistent with those of Swayne et al. (9). The inability of the virus to replicate efficiently in chickens and ducks could very well be linked to its human virus–like receptor recognition.
The ability of influenza A viruses to agglutinate erythrocytes from a variety of hosts may reflect the viruses’ receptor specificity (10,11). We observed similar binding patterns for the mammalian influenza (H1N1) viruses, with the exception that the pandemic strains had reduced binding to chicken erythrocytes. This binding pattern was also observed with 1 of the swine isolates, suggesting it might be a trait of swine-adapted viruses. Taken together, a difference in the hemadsorption phenotype observed with erythrocytes from species with either less Neu5Acα2,6Gal or less Neu5Ac linkage overall could be explained by the mammalian origin of the novel pandemic (H1N1) 2009 influenza viruses.
To test this possibility, we measured the HA affinity of H1 influenza viruses from various species of origin for synthetic receptor analogues. All mammalian H1 viruses showed a typical human virus–like preference for the Neu5Acα2,6Gal-containing receptor 6′SLN. Compared with the currently circulating H1N1 human viruses, both pandemic (H1N1) 2009 strains and contemporary swine influenza virus (H1N1) strains were able to bind substantially more strongly (5–12×) to an α2,6-containing glycopolymer; the currently circulating subtype H1N1 human viruses are strictly adapted to this receptor (12). This feature demonstrated that pandemic H1N1 strains, which have a HA gene of swine lineage, have retained their current swine virus–like binding characteristics despite their efficient spread in humans.
To identify substitutions in the HA molecule that could be responsible for the human-like receptor binding phenotype of the pandemic and contemporary swine influenza (H1N1) isolates, we compared the H1 HA sequences deposited in the Influenza Research Database (www.fludb.org). We observed that 99.99% of all swine viruses isolated after 1980 have Asp190 or Asn190. HA sequences of swine viruses isolated before 2000 harbor Gly225, whereas 92.3% of more recent classical swine viruses have Asp225. Crystallographic analysis of human and swine H1 HA has shown that Asp225 and Asp190 are responsible for human virus–like receptor specificity (13). Therefore, the human-like amino acids encoded at HA positions 190 and 225 in the novel pandemic and swine influenza (H1N1) viruses may at least partially explain their innate affinity for the human-type receptor.
Recent phylogenetic analysis showed that each segment of the pandemic (H1N1) 2009 virus is nested within a well-established swine influenza lineage for >10 years before the recent outbreak (7). Hence, the ancestors of this virus circulated undetected for about a decade before the virus emerged in humans. Our finding that contemporary swine viruses acquired the ability to recognize 6′SLN with at least 5-fold higher affinity than did human strains and completely lost the ability to bind to Neu5Acα2,3Gal provides clear evidence to support this hypothesis. It is possible that the progenitors of pandemic (H1N1) 2009 virus were accumulating enough mammal-associated changes to allow a refinement of their receptor-binding properties. Our findings substantiate that strong mammalian-like receptor specificity is a critical barrier to infection of various hosts with pandemic (H1N1) 2009 virus. Other biological factors associated with their adaptation and tissue tropism in humans will likely be identified in the future.
Dr Ilyushina is a postdoctoral research fellow in the Division of Virology, Department of Infectious Diseases, St. Jude Children’s Research Hospital, Memphis, Tennessee, USA. She has research interests in the molecular basis of viral pathogenesis and antiviral drug usage for the control of influenza infection, including emerging influenza (H1N1) and (H5N1) viruses.
Acknowledgments
We thank Marie R. Gramer for provision of A/swine/North Carolina/007270/2008 (H1N1) and A/swine/Iowa/003479/2009 (H1N1) influenza A virus strains; Elena A. Govorkova for helpful discussion; Jon P. Seiler for technical assistance; Betsy Williford for illustrations; and David Galloway and Sharon Naron for editorial assistance.
This research was funded through contract no. HHSN266200700005C from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services; a TBP grant from the Korea Research Council of Fundamental Science & Technology; grant no. KGM3110912 from the Korea Research Institute of Bioscience and Biotechnology Research Initiative Program of Korea; a Presidium Grant “Molecular and Cell Biology” from the Russian Academy of Sciences; and the American Lebanese Syrian Associated Charities.
References
- Centers for Disease Control and Prevention. Swine influenza A (H1N1) infection in two children—southern California, March–April 2009. MMWR Morb Mortal Wkly Rep. 2009;58:400–2.PubMedGoogle Scholar
- World Health Organization. Global Alert and Response. Pandemic (H1N1) 2009–update 58. 2009 Jul 6 [cited 2009 Jul 9]. http://www.who.int/csr/don/2009_07_06/en/index.html
- Nicholls JM, Chan RWY, Russell RJ, Air GM, Peiris JSM. Evolving complexities of influenza virus and its receptors. Trends Microbiol. 2008;16:149–57. DOIPubMedGoogle Scholar
- Stevens J, Blixt O, Glaser L, Taubenberger JK, Palese P, Paulson JC, Glycan microarray analysis of the hemagglutinins from modern and pandemic influenza viruses reveals different receptor specificities. J Mol Biol. 2006;355:1143–55. DOIPubMedGoogle Scholar
- Matrosovich M, Tuzikov A, Bovin N, Gambaryan AS, Klimov A, Castrucci MR, Early alterations of the receptor-binding properties of H1, H2, and H3 avian influenza virus hemagglutinins after their introduction into mammals. J Virol. 2000;74:8502–12. DOIPubMedGoogle Scholar
- Garten RJ, Davis CD, Russell CA, Shu B, Lindstrom S, Balish A, Antigenic and genetic characteristics of swine-origin 2009 A (H1N1) influenza viruses circulating in humans. Science. 2009;325:197–201. DOIPubMedGoogle Scholar
- Smith GJD, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature. 2009;459:1122–5. DOIPubMedGoogle Scholar
- Reed LJ, Muench H. A simple method for estimating fifty percent endpoints. Am J Hyg. 1938;27:493–7.
- Swayne DE, Pantin-Jackwood M, Kapczynski D, Spackman E, Suarez DL. Susceptibility of poultry to pandemic (H1N1) 2009 virus [letter]. Emerg Infect Dis [serial on the Internet]. 2009 Dec [cited 2009 Nov 18]. http://www.cdc.gov/EID/content/15/12/2061.htm
- Ito T, Suzuki Y, Mithaul L, Vines A, Kida H, Kawaoka Y. Receptor specificity of influenza A viruses correlates with the agglutination of erythrocytes from different animal species. Virology. 1997;227:493–9. DOIPubMedGoogle Scholar
- Medeiros R, Escriou N, Naffakh N, Manuguerra JC, van der Werf S. Hemagglutinin residues of recent human A (H3N2) influenza viruses that contribute to the inability to agglutinate chicken erythrocytes. Virology. 2001;289:74–85. DOIPubMedGoogle Scholar
- Mochalova L, Gambaryan A, Romanova J, Tuzikov A, Chinarev A, Katinger D, Receptor-binding properties of modern human influenza viruses primarily isolated in Vero and MDCK cells and chicken embryonated eggs. Virology. 2003;313:473–80. DOIPubMedGoogle Scholar
- Gamblin SJ, Haire LF, Russell RJ, Stevens DJ, Xiao B, Ha Y, The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science. 2004;303:1838–42. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 16, Number 2—February 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Robert G. Webster, Department of Infectious Diseases, St. Jude Children’s Research Hospital, 262 Danny Thomas Pl, Memphis, TN 38105-3678, USA
Top