Volume 19, Number 6—June 2013
Synopsis
New Delhi Metallo-β-Lactamase–producing Enterobacteriaceae, United States
Susceptibility to Selected Antimicrobial Agents
BMD Screening for Metallo-β-Lactamase
Modified Hodge Test
Etest for Detection of MBLs
Detection of blaNDM and blaKPC by Real-Time PCR
DNA Sequence Analysis of blaNDM
Plasmid Isolation and Transformation
Characterization of blaNDM-bearing Plasmids
Pulsed-Field Gel Electrophoresis
Multilocus Sequence Typing
Clinical and Epidemiologic Information
Antimicrobial Drug Susceptibility Patterns
Detection of NDM Producers
Sequencing of the blaNDM Gene
Antimicrobial Drug Resistance Transferred on the blaNDM Plasmid
blaNDM-bearing Plasmids
Strain Typing
Conclusions
Cite This Article
Cite This Article
Citation for Media
Abstract
We characterized 9 New Delhi metallo-β-lactamase–producing Enterobacteriaceae (5 Klebsiella pneumoniae, 2 Escherichia coli, 1 Enterobacter cloacae, 1 Salmonella enterica serovar Senftenberg) isolates identified in the United States and cultured from 8 patients in 5 states during April 2009–March 2011. Isolates were resistant to β-lactams, fluoroquinolones, and aminoglycosides, demonstrated MICs ≤1 µg/mL of colistin and polymyxin, and yielded positive metallo-β-lactamase screening results. Eight isolates had blaNDM-1, and 1 isolate had a novel allele (blaNDM-6). All 8 patients had recently been in India or Pakistan, where 6 received inpatient health care. Plasmids carrying blaNDM frequently carried AmpC or extended spectrum β-lactamase genes. Two K. pneumoniae isolates and a K. pneumoniae isolate from Sweden shared incompatibility group A/C plasmids with indistinguishable restriction patterns and a common blaNDM fragment; all 3 were multilocus sequence type 14. Restriction profiles of the remaining New Delhi metallo-β-lactamase plasmids, including 2 from the same patient, were diverse.
During the past decade, there has been an emergence of carbapenem-resistant Enterobacteriaceae that produce carbapenemases, enzymes that efficiently hydrolyze carbapenems, as well as most β-lactam drugs (1). The most common carbapenemase among Enterobacteriaceae in the United States is the Ambler class A Klebsiella pneumoniae carbapenemase (KPC), an enzyme that is found throughout the United States and globally (2,3). The emergence of another group of carbapenemases, the Ambler class B metallo-β-lactamases (MBLs), is of great concern worldwide (4). Until recently, MBLs were rarely identified in the United States and were found exclusively in Pseudomonas aeruginosa (5). However, recent reports of producing IMP– and VIM-type MBLs K. pneumoniae (6,7) have increased concerns over additional transmissible carbapenem resistance mechanisms in Enterobacteriaceae.
Among the most recent carbapenemases to appear in the United States is the newly described New Delhi MBL (NDM) (8–12). First reported in 2009, NDM-1 was initially identified in K. pneumoniae and Escherichia coli clinical isolates obtained from a Swedish patient who had been hospitalized in India (13). Drug-resistant gram-negative bacteria that produce NDM have been found in community and health care settings in India and Pakistan in a wide range of gram-negative genera containing diverse blaNDM-harboring plasmids, and have been reported in >15 countries worldwide (4,14,15). The widespread dissemination of NDM-producing isolates and the apparent ease of mobility of blaNDM is a major threat to public health on a global scale.
To complement reports of individual cases (8,10,12), we performed extensive laboratory characterization of 9 clinical isolates of NDM-producing Enterobacteriaceae collected from patients in the United States during April 2009–March 2011. Strain typing and plasmid restriction analysis were performed to identify common lineages. We also describe a novel NDM-encoding allele, designated blaNDM-6.
Nine clinical isolates (5 K. pneumoniae, 2 E. coli, 1 Enterobacter cloacae, and 1 Salmonella enterica serovar Senftenberg), were collected from 8 patients during April 2009–March 2011 and submitted to the Centers for Disease Control and Prevention (Atlanta, GA, USA) for reference susceptibility testing during January 2010–April 2011 (Table 1). Four patients were from California and 1 each was from Illinois, Maryland, Massachusetts, and Virginia. Species identification was confirmed with the Vitek 2 automated system (bioMérieux Vitek Systems Inc., Hazelwood, MO, USA). The S. enterica serovar Senftenberg isolate was further classified by serotyping (12). A previously identified NDM-1–producing K. pneumoniae isolate (0S-506) from Sweden (13) was used as a positive control for phenotypic and molecular characterization methods, including strain typing of K. pneumoniae isolates. As part of a public health intervention for each of these isolates, an epidemiologist from CDC contacted local health departments and providers to identify characteristics of patients from whom NDM-producing isolates were obtained.
MICs of amikacin, aztreonam, cefotaxime, cefepime, ciprofloxacin, colistin, doripenem, ertapenem, gentamicin, imipenem (IMP), meropenem, polymyxin B, tetracycline, tigecycline, and trimethoprim/sulfamethoxazole were determined by using reference broth microdilution (BMD) with panels prepared in-house according to Clinical and Laboratory Standards Institute (Wayne, PA, USA) guidelines (16) and stored at −70°C until use. MICs of tigecycline were interpreted according to breakpoints established by the US Food and Drug Administration (Silver Spring, MD, USA) (www.rxlist.com/tygacil-drug.htm). E. coli ATCC 25922, K. pneumoniae ATCC BAA-2146, P. aeruginosa ATCC 27853, Staphylococcus aureus ATCC 29213, and Enterococcus faecalis ATCC 29212 were used for quality control.
MICs in the absence and presence of a combination of 0.2 mmol/L EDTA (Sigma-Aldrich, St. Louis, MO, USA) and 0.02 mmol/L 1,10-phenanthroline (Acros Organics, Geel, Belgium) were determined as described (17) by using screening wells containing IMP at concentrations ranging from 0.25 µg/mL through 1,024 µg/mL. A ratio ≥4 in the IMP MIC compared with the IMP MIC in the presence of chelators (IMP+EP) was considered a positive result for MBL production. K. pneumoniae ATCC BAA-2146 and P. aeruginosa ATCC 27853 were used as positive and negative controls, respectively.
The Modified Hodge test (MHT), which is recommended by Clinical and Laboratory Standards Institute as a confirmatory test for carbapenemase production (16), was performed for each strain with 10-µg disks containing meropenem and ertapenem (Becton-Dickinson, Sparks, MD, USA). K. pneumoniae ATCC BAA-1705 and BAA-1706 were used as positive and negative controls, respectively.
Detection of MBLs was performed with Etest MBL strips (AB bioMérieux, St. Louis, MO, USA) containing IMP (IP) and IMP + EDTA (IPI). Strips were used according to instructions provided by the manufacturer.
A multiplexed Taqman-based real-time PCR for blaNDM and blaKPC, as well as the universal bacterial 16S rRNA–encoding gene (18) as an endogenous control for DNA amplification, was performed on the 7500 Fast system (Applied Biosystems, Carlsbad, CA, USA). Cell lysates were prepared as described (19). Each PCR (20-μL volume) included 1× QuantiFast Probe PCR Master Mix (QIAGEN, Valencia, CA, USA), a combined primer/probe solution with final concentrations of 500 nmol/L for each primer and 250 nmol/L for each probe (Table 2), and 2 μL of template. Included in each assay were a blaNDM-positive control (K. pneumoniae ATCC BAA-2146), a blaKPC-positive control (K. pneumoniae ATCC BAA-1705), a carbapenemase-negative control (K. pneumoniae ATCC BAA-1706), and a no template control. Cycling conditions were a 3-min enzyme activation step at 95°C, followed by 40 cycles for 3 s at 95°C and 30 s at 60°C.
Reactions with 16S cycle threshold (Ct) values of 10–30 were considered valid, those with NDM or KPC Ct values of 10–30 were considered NDM positive or KPC negative, and those with NDM or KPC Ct values ≥40 were considered NDM positive or KPC negative (www.cdc.gov/HAI/settings/lab/kpc-ndm1-lab-protocol.html).
Forward and reverse primers outside the blaNDM coding region (Table 2) were used to amplify a 1,013-bp product. Bidirectional DNA sequencing of blaNDM was determined from independent products with primers used for amplification, as well as blaNDM internal primers (Table 2).
Plasmid DNA was isolated from 50-mL overnight cultures by using a Plasmid Midi Kit (QIAGEN) according to the manufacturer’s protocol. To enhance the yield of large, low-copy plasmids, DNA was eluted with elution buffer prewarmed to 65°C. E. coli DH10BT1 competent cells (Invitrogen, Carlsbad, CA, USA) were transformed with plasmid DNA by electroporation (Gene Pulser Xcell; Bio-Rad Laboratories, Hercules, CA, USA) according to the manufacturer’s instructions. Transformants were selected on Luria-Bertani agar containing 1 µg/mL of meropenem and were screened by real-time PCR for blaNDM. Transformant plasmid DNA was evaluated by electrophoresis, and a representative transformant containing a single NDM-encoding plasmid was chosen for further study (designated by TF suffix). E. coli NCTC50192, which contained 4 plasmids (≈154, 66, 38, and 7 kb) (20), and E. coli V517, which contained 8 plasmids ranging from ≈56.4 kb to 2.2 kb (21), were used as plasmid size standards.
Plasmid DNA from each transformant was digested with XmnI (New England Biolabs, Ipswich, MA, USA), separated by electrophoresis, transferred to a nylon membrane (Zeta-Probe; Bio-Rad Laboratories), and hybridized with an 808-bp digoxigenin (DIG)–labeled blaNDM probe (Table 2) by using the PCR DIG Probe Synthesis Kit (Roche Applied Science, Mannheim, Germany). Hybridization at 42°C, washes, and detection by using a DIG Luminescent Detection Kit (Roche Applied Science) were performed according to the manufacturer’s instructions.
NDM-encoding plasmids were assigned to an incompatibility group by using PCR replicon typing as described (22). Additional β-lactamases that were co-transferred with each blaNDM-carrying plasmid were identified by using the Check-MDR CT101 Microarray Assay (Check-Points BV, Wageningen, the Netherlands), which detects genes encoding extended-spectrum β-lactamases (ESBLs) (TEM, SHV, and CTX-M), plasmid-mediated AmpCs (CMY, DHA, FOX, MOX, ACC, MIR, and ACT), as well as KPC and NDM (23). PCR was used to screen for armA and rmtC 16S rRNA methylase genes that confer resistance to aminoglycosides (24).
Pulsed-field gel electrophoresis (PFGE) was performed by using the CHEF mapper electrophoresis system (Bio-Rad Laboratories) with XbaI-digested K. pneumoniae and E. coli chromosomal DNA, as described for E. coli (www.cdc.gov/pulsenet/protocols.htm). PFGE patterns were compared by using the Dice coefficient and clustering by using the unweighted-pair group method with average linkages (Bionumerics version 5.10; Applied Maths Inc., Austin, TX, USA).
Multilocus sequence typing (MLST) was used to classify K. pneumoniae and E. coli isolates. This procedure was performed and results were interpreted according to protocols on the Institut Pasteur MLST database website (www.pasteur.fr/recherche/genopole/PF8/mlst) (25,26).
We identified blaNDM by real-time PCR and DNA sequence analysis for 9 clinical isolates received at the Centers for Disease Control and Prevention during January 2010–April 2011 from 8 patients (Table 1) in 5 states. Two isolates, K. pneumoniae 1100975 and S. enterica serovar Senftenberg 1101168, were isolated from the same patient 1 month apart from a clinical specimen and a surveillance specimen, respectively (12). NDM-producing Enterobacteriaceae were isolated from a variety of specimen sources, including urine (4/9), respiratory samples (3/9), feces (1/9), and blood (1/9) (Table 1), and mostly represented colonization. All 8 patients (age range 13 months–73 years, median 62.5 years) had a recent travel history (within 4 months) that included India or Pakistan, during which 6 patients received inpatient medical care, and 1 received outpatient medical care. One patient was a citizen of India who traveled frequently between the United States and India. All medical exposures abroad resulted from medical problems that occurred in that country; none of the patients had traveled for the purpose of obtaining medical care (i.e., medical tourism).
All 9 NDM-producing isolates from the United States and K. pneumoniae 0S-506 from Sweden were resistant to all β-lactams tested (including carbapenems and aztreonam), ciprofloxacin, amikacin, and gentamicin, and demonstrated MICs ≤1 μg/mL for colistin and polymyxin B; 7/9 were susceptible (MIC ≤2 μg/mL) to tigecycline. Only 2 isolates were susceptible to tetracycline, and only the S. enterica serovar Senftenberg isolate was susceptible to trimethoprim/sulfamethoxazole (Table 3).
Although each of the 9 isolates showed resistance to carbapenems, detection of carbapenemase activity by using the MHT was variable (Table 3). Six of 9 isolates had a positive MHT result for meropenem and ertapenem, and 3 were positive for ertapenem but negative for meropenem. K. pneumoniae 0S-506 was MHT negative for both carbapenems. The Etest MBL result was positive for K. pneumoniae 0S-506 and for 6/9 isolates from the United States (IP:IPI ratio ≥12). The remaining 3 isolates showed either a phantom zone or deformed ellipse, which are also indicative of an MBL according to the AB Biodisk information, although deformation of the ellipse can be difficult to recognize (10). The BMD MBL screen provided the most conclusive results for MBL detection: all 9 isolates and K. pneumoniae 0S-506 demonstrated an MIC IMP:IMP+EP ratio ≥8, which is indicative of MBL production (Table 3).
DNA sequencing of blaNDM from each of the 9 isolates showed that 8 encoded NDM-1, but the coding sequence in E. coli 1100101 and its transformant differed from that of blaNDM-1 (GenBank accession no. FN396876) by a C→T modification at nucleotide position 698, resulting in an alanine→valine substitution at aa 233 in the inferred protein. This novel NDM variant has been designated NDM-6 (G.A. Jacoby and K. Bush, www.lahey.org/Studies) and its sequence has been deposited in GenBank under accession no. JN967644.
Because the NDM-producing isolates from the United States and K. pneumoniae 0S-506 from Sweden contained multiple plasmids (range 3–6 plasmids), the blaNDM-bearing plasmid from each isolate was transferred to a plasmid-negative E. coli by electroporation. A transformant carrying only the blaNDM-bearing plasmid was chosen for further study.
Eight transformants were either resistant or intermediate to cefepime, cefotaxime, and all carbapenems tested, but transformant 1100192-TF was susceptible to meropenem (Table 3). Five transformants remained resistant to aztreonam, indicating that an additional resistance mechanism (e.g., AmpC or ESBL) was also carried on the NDM-encoding plasmid because MBLs do not independently hydrolyze aztreonam. Resistance to amikacin and gentamicin was co-transferred in 7 instances, but 1100192-TF remained susceptible to both drugs, and 1100101-TF was resistant only to amikacin. All transformants remained susceptible to ciprofloxacin and tetracycline, and only 1000654-TF showed resistance to trimethoprim/sulfamethoxazole.
The MHT identified carbapenemase activity in all transformant strains, although 3 were positive with only 1 of 2 carbapenems (Table 3). Only transformant 1000654-TF showed an Etest MBL IP:IPI ratio ≥8; the remainder had either a phantom zone or an ellipse deformation. The BMD MBL screening method detected MBL production in all transformants (Table 3).
Additional β-lactamase genes carried in parental strains and transformants were identified by using the Check-MDR CT101 microarray assay (23) (Table 4). Three transformants that remained susceptible to aztreonam had only the NDM β-lactamase. Transformants with aztreonam resistance carried either a CMY-II-type AmpC (n = 4) or a CTX-M-1-type ESBL (n = 2), in addition to blaNDM.
Four of 8 transformants resistant to amikacin and gentamicin contained armA, and 2 contained rmtC (Table 4), both of which are 16S rRNA methylase genes that confer high-level resistance to nearly all aminoglycosides (24). The mechanism conferring aminoglycoside resistance in the remaining 2 transformants was not caused by armA or rmtC (Table 4) and was not characterized further.
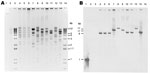
The incompatibility groups of NDM-encoding plasmids were primarily A/C (n = 4), but also included FII (n = 2), L/M (n = 1) and 2 plasmids that were untypeable (Table 4). Eight XmnI restriction patterns were observed among the NDM-encoding plasmids isolated from transformants of the isolates from the United States and isolate 0S-506 from Sweden (Figure 1). Plasmid restriction profiles from K. pneumoniae 0S-506 and 2 K. pneumoniae isolates (1100770 and 1100975) were indistinguishable. Each isolate carried blaNDM on an XmnI fragment of ≈6 kb (Figure 1) and had similar transferred antimicrobial susceptibility profiles (Table 3); carried the same ESBL and AmpC genes; and had plasmid replicon type A/C (Table 4). Other blaNDM-bearing plasmids were diverse, including those isolated from the same patient (K. pneumoniae 1100975 and S. enterica serovar senftenberg 1101168) (Figure 1).
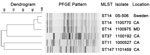
The K. pneumoniae isolates with indistinguishable blaNDM plasmid profiles were closely related by PFGE, and all were classified as sequence type (ST)14 by MLST (Figure 2) (13). The remaining K. pneumoniae (Figure 2) and E. coli (data not shown) isolates showed more diverse PFGE patterns and MLST types, including ST37, ST11, and ST147. E. coli isolates were identified as ST500 and ST43. For most isolates, ST43 corresponds to ST131 in the MLST scheme of Wirth et al. (27) (S. Brisse, pers. comm.).
The 9 NDM-producing isolates described were resistant to all β-lactams, including aztreonam, as well as all commonly used aminoglycosides and fluoroquinolones. In addition to NDM, each isolate carried >1 other β-lactamase, including CMY-II-type AmpCs and/or CTX-M-1-type ESBLs (which co-transferred with NDM for all but 3 isolates). Most blaNDM-bearing plasmids also carried armA or rmtC 16S rRNA methylase genes, which confer high-level resistance to most aminoglycosides and are often associated with these plasmids (24,28). Although resistance to ciprofloxacin and tetracycline did not transfer with the blaNDM-bearing plasmid, trimethoprim/sulfamethoxazole resistance was conferred to 1 transformant. For several strains, the transformant displayed decreased carbapenem resistance compared with a parental strain (e.g., 1100192-TF), suggesting that additional mechanisms (e.g., AmpC and ESBL) present in the parental strain and not carried on the NDM-encoding plasmid may have contributed to the initial carbapenem-resistant Enterobacteriaceae phenotype observed. These findings emphasize the diversity of resistance mechanisms carried on NDM-encoding plasmids, as reported (28).
We used 3 screening methods for phenotypic detection of MBL activity: MHT, Etest MBL, and BMD MBL. The MHT was not sensitive for detection of NDM activity; 3 isolates were positive only for 1 carbapenem tested, and K. pneumoniae 0S-506, the first characterized NDM-producing isolate (13), was negative for both carbapenems. Etest MBL definitively identified 6 parental isolates and 1 NDM-producing transformant as MBL producers, but 3 parental strains and 8 transformants displayed a phantom zone or slight deformation of the IP or IPI ellipse. The Etest MBL package insert states that these findings are indicative of an MBL, but the ellipse deformation we observed was subtle and less dramatic than the example provided. The BMD MBL screen provided the most unambiguous results, and yielded IMP to IMP + EP MIC ratios ≥8 for all NDM-producing parental and transformant strains. In an earlier validation study, this BMD MBL screen had a sensitivity of 95% and a specificity of 100% (29).
We reliably detected blaNDM with a novel multiplexed real-time PCR designed to detect the blaNDM and blaKPC genes. DNA sequence analysis confirmed the PCR results and identified the blaNDM allele in each isolate. One isolate contained a variant allele designated blaNDM-6. An NDM-6–producing E. coli strain was also recently identified in a patient in New Zealand who had received medical care in India (30).
Plasmids carrying blaNDM have been reported to range in size from 50 through 400 kb (14,15). Because all isolates in this report carried multiple plasmids, it was necessary to transfer the NDM-encoding plasmid to a plasmid-negative recipient for analysis. Three K. pneumoniae isolates, including the original NDM-producer from Sweden (13), contained an ≈170-kb blaNDM–bearing plasmid, and each isolate was indistinguishable by restriction analysis and Southern blot. These strains were also closely related by PFGE and MLST (ST14). Furthermore, the antimicrobial drug susceptibility profiles of their parental and transformant isolates were similar. In contrast, the remaining isolates contained different blaNDM-bearing plasmids ranging in size from 100 kb through 200 kb, carried blaNDM on different restriction fragments, and were not related by PFGE or MLST. Most of the blaNDM-bearing plasmids belonged to incompatibility groups A/C or L/M, both broad host range plasmids, and FII, a narrow host range plasmid (31). All 3 replicon types have been found to be associated with a variety of β-lactam resistance mechanisms in Enterobacteriaceae (32). These findings were consistent with reports of extensive diversity among blaNDM-bearing plasmids in Enterobacteriaceae (14).
The MLST types identified among NDM-producing K. pneumoniae described here have been associated with various resistant strains of K. pneumoniae worldwide (28,33). ST11, ST147, and ST15, a single locus variant of ST14, have been identified as epidemic clones of CTX-M-15–producing K. pneumoniae in Hungary (34). In addition, ST11 is the dominant KPC-producing strain in China (35) and is a single locus variant of ST258, the dominant KPC-producing strain in the United States (2). ST11 NDM-producing K. pneumoniae strains were among the first NDM-producing Enterobacteriaceae reported in New Zealand (30), and ST147 NDM-producing K. pneumoniae isolates have been reported in Switzerland (28), Canada (36), Australia (37), and in an Iraqi patient transferred to a hospital in France (28). ST14 has been identified among KPC-producing strains in the United States (38), and is associated with NDM-producing K. pneumoniae isolates in Kenya and the Sultanate of Oman (28), and as the most frequently encountered ST in a recent study of NDM-1–producing K. pneumoniae from 3 countries (33). We also report a ST43/ST131 NDM-producing E. coli strain in our study. This clone is most notably associated with the global dissemination of the CTX-M-15 ESBL (39).
Each patient associated with the isolates described here had recently been in India or Pakistan, and most had received inpatient medical care in those countries. The link between NDM acquisition and health care exposure abroad has been extensively described (4,15,40). However, 1 patient only had outpatient health care during travel, and another had no documented health care, although the second patient had several active medical problems, including the presence of an invasive medical device during travel. In contrast to the early NDM case-patients reported in the United Kingdom (15), none of the patients in our study had traveled specifically for the purpose of obtaining medical care.
Several factors contribute to the global dissemination of blaNDM as it spreads through a variety of plasmids and bacterial strains. The environmental and epidemiologic factors driving this spread and the molecular mechanisms by which it disseminates are not well understood. However, in the 3 years since its initial description, NDM has spread rapidly worldwide and has now been described in >15 countries in 5 continents (4,8). Since the completion of this study, numerous additional NDM-producing Enterobacteriaceae have been identified in the United States. The relative ease with which this resistance mechanism appears to move within and between different bacterial genera, as well as mobility of humans infected or colonized with NDM producers, serves to highlight the need for reliable and rapid means of detecting drug-resistant organisms to implement infection control measures to prevent further dissemination.
Dr. Rasheed is team lead of the Antimicrobial Resistance and Characterization Laboratory, Division of Healthcare Quality Promotion, at the Centers for Disease Control and Prevention in Atlanta. His primary research interest is molecular characterization of antimicrobial drug resistance mechanisms involved with healthcare-associated pathogens.
Acknowledgment
We thank Christian Giske for providing K. pneumoniae isolate 0S-506, Maria Karlsson for providing armA- and rmtC-positive control strains, and Alessandra Carattoli for providing control strains for PCR replicon typing.
References
- Queenan AM, Bush K. Carbapenemases: the versatile β-lactamases. Clin Microbiol Rev. 2007;20:440–58. DOIPubMedGoogle Scholar
- Kitchel B, Rasheed JK, Patel JB, Srinivasan A, Navon-Venezia S, Carmeli Y, Molecular epidemiology of KPC-producing Klebsiella pneumoniae isolates in the United States: clonal expansion of multilocus sequence type 258. Antimicrob Agents Chemother. 2009;53:3365–70. DOIPubMedGoogle Scholar
- Nordmann P, Cuzon G, Naas T. The real threat of Klebsiella pneumoniae carbapenemase-producing bacteria. Lancet Infect Dis. 2009;9:228–36. DOIPubMedGoogle Scholar
- Nordmann P, Naas T, Poirel L. Global spread of carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis. 2011;17:1791–8. DOIPubMedGoogle Scholar
- Hanson ND, Hossain A, Buck L, Moland ES, Thomson KS. First occurrence of a Pseudomonas aeruginosa isolate in the United States producing an IMP metallo-β-lactamase, IMP-18. Antimicrob Agents Chemother. 2006;50:2272–3. DOIPubMedGoogle Scholar
- Limbago BM, Rasheed JK, Anderson KF, Zhu W, Kitchel B, Watz N, IMP-producing carbapenem-resistant K. pneumoniae in the United States. J Clin Microbiol. 2011;49:4239–45. DOIPubMedGoogle Scholar
- Gupta N, Limbago BM, Patel JB, Kallen AJ. Carbapenem-resistant Enterobacteriaceae: epidemiology and prevention. Clin Infect Dis. 2011;53:60–7. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Detection of Enterobacteriaceae isolates carrying metallo-beta-lactamase—United States, 2010. MMWR Morb Mortal Wkly Rep. 2010;59:750 .PubMedGoogle Scholar
- Peirano G, Schreckenberger PC, Pitout JD. Characteristics of NDM-1-producing Escherichia coli isolates that belong to the successful and virulent clone ST131. Antimicrob Agents Chemother. 2011;55:2986–8. DOIPubMedGoogle Scholar
- Mochon AB, Garner OB, Hindler JA, Krogstad P, Ward KW, Lewinski MA, New Delhi metallo-β-lactamase (NDM-1)–producing Klebsiella pneumoniae: case report and laboratory detection strategies. J Clin Microbiol. 2011;49:1667–70. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Carbapenem-resistant Enterobacteriaceae containing New Delhi metallo-beta-lactamase in two patients—Rhode Island, March 2012. MMWR Morb Mortal Wkly Rep. 2012;61:446–8 .PubMedGoogle Scholar
- Savard P, Gopinath R, Zhu W, Kitchel B, Rasheed JK, Tekle T, The first NDM-positive Salmonella spp. identified in the United States. Antimicrob Agents Chemother. 2011;55:5957–8. DOIPubMedGoogle Scholar
- Yong D, Toleman MA, Giske CG, Cho HS, Sundman K, Lee K, Characterization of a new metallo-β-lactamase gene, blaNDM-1, and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob Agents Chemother. 2009;53:5046–54. DOIPubMedGoogle Scholar
- Walsh TR, Weeks J, Livermore DM, Toleman MA. Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study. Lancet Infect Dis. 2011;11:355–62. DOIPubMedGoogle Scholar
- Kumarasamy KK, Toleman MA, Walsh TR, Bagaria J, Butt F, Balakrishnan R, Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis. 2010;10:597–602. DOIPubMedGoogle Scholar
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Twenty-first informational supplement; CLSI document M100–S21. Wayne (PA): The Institute; 2011.
- Migliavacca R, Docquier JD, Mugnaioli C, Amicosante G, Daturi R, Lee K, Simple microdilution test for detection of metallo-β-lactamase production in Pseudomonas aeruginosa. J Clin Microbiol. 2002;40:4388–90. DOIPubMedGoogle Scholar
- Yang S, Lin S, Kelen GD, Quinn TC, Dick JD, Gaydos CA, Quantitative multiprobe PCR assay for simultaneous detection and identification to species level of bacterial pathogens. J Clin Microbiol. 2002;40:3449–54. DOIPubMedGoogle Scholar
- Kitchel B, Rasheed JK, Endimiani A, Hujer AM, Anderson KF, Bonomo RA, Genetic factors associated with elevated carbapenem resistance in KPC-producing Klebsiella pneumoniae. Antimicrob Agents Chemother. 2010;54:4201–7. DOIPubMedGoogle Scholar
- Macrina FL, Kopecko DJ, Jones KR, Ayers DJ, McCowen SM. A multiple plasmid-containing Escherichia coli strain: convenient source of size reference plasmid molecules. Plasmid. 1978;1:417–20. DOIPubMedGoogle Scholar
- Johnson TJ, Wannemuehler YM, Johnson SJ, Logue CM, White DG, Doetkott C, Plasmid replicon typing of commensal and pathogenic Escherichia coli isolates. Appl Environ Microbiol. 2007;73:1976–83. DOIPubMedGoogle Scholar
- Bogaerts P, Hujer AM, Naas T, de Castro RR, Endimiani A, Nordmann P, Multicenter evaluation of a new DNA microarray for rapid detection of clinically relevant bla genes from β-lactam-resistant gram-negative bacteria. Antimicrob Agents Chemother. 2011;55:4457–60. DOIPubMedGoogle Scholar
- Berçot B, Poirel L, Nordmann P. Updated multiplex polymerase chain reaction for detection of 16S rRNA methylases: high prevalence among NDM-1 producers. Diagn Microbiol Infect Dis. 2011;71:442–5. DOIPubMedGoogle Scholar
- Diancourt L, Passet V, Verhoef J, Grimont PA, Brisse S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J Clin Microbiol. 2005;43:4178–82. DOIPubMedGoogle Scholar
- Jaureguy F, Landraud L, Passet V, Diancourt L, Frapy E, Guigon G, Phylogenetic and genomic diversity of human bacteremic Escherichia coli strains. BMC Genomics. 2008;9:560. DOIPubMedGoogle Scholar
- Wirth T, Falush D, Lan R, Colles F, Mensa P, Wieler LH, Sex and virulence in Escherichia coli: an evolutionary perspective. Mol Microbiol. 2006;60:1136–51. DOIPubMedGoogle Scholar
- Poirel L, Dortet L, Bernabeu S, Nordmann P. Genetic features of blaNDM-1-positive Enterobacteriaceae. Antimicrob Agents Chemother. 2011;55:5403–7. DOIPubMedGoogle Scholar
- Anderson KF, Rasheed JK, Kitchel B, Wong B, Clark N, Limbago B. Validation of a broth screen for the detection of metallo-β-lactamase. In: Abstracts of the 111th General Meeting, American Society of Microbiology, New Orleans, May 21–24, 2011. Washington (DC): American Society for Microbiology Press; 2011. Abstract no. C-611, p. 104.
- Williamson DA, Sidjabat HE, Freeman JT, Roberts SA, Silvey A, Woodhouse R, Identification and molecular characterisation of New Delhi metallo-β-lactamase-1 (NDM-1)– and NDM-6–producing Enterobacteriaceae from New Zealand hospitals. Int J Antimicrob Agents. 2012;39:529–33 . DOIPubMedGoogle Scholar
- Novais A, Canton R, Moreira R, Peixe L, Baquero F, Coque TM. Emergence and dissemination of Enterobacteriaceae isolates producing CTX-M-1–like enzymes in Spain are associated with IncFII (CTX-M-15) and broad-host-range (CTX-M-1, -3, and -32) plasmids. Antimicrob Agents Chemother. 2007;51:796–9 . DOIPubMedGoogle Scholar
- Carattoli A. Resistance plasmid families in Enterobacteriaceae. Antimicrob Agents Chemother. 2009;53:2227–38. DOIPubMedGoogle Scholar
- Giske CG, Froding I, Hasan CM, Turlej-Rogacka A, Toleman M, Livermore D, Diverse sequence types of Klebsiella pneumoniae contribute to the dissemination of blaNDM-1 in India, Sweden, and the United Kingdom. Antimicrob Agents Chemother. 2012;56:2735–8. DOIPubMedGoogle Scholar
- Damjanova I, Toth A, Paszti J, Hajbel-Vekony G, Jakab M, Berta J, Expansion and countrywide dissemination of ST11, ST15 and ST147 ciprofloxacin-resistant CTX-M-15-type β-lactamase-producing Klebsiella pneumoniae epidemic clones in Hungary in 2005–the new ‘MRSAs’? J Antimicrob Chemother. 2008;62:978–85. DOIPubMedGoogle Scholar
- Qi Y, Wei Z, Ji S, Du X, Shen P, Yu Y. ST11, the dominant clone of KPC-producing Klebsiella pneumoniae in China. J Antimicrob Chemother. 2011;66:307–12. DOIPubMedGoogle Scholar
- Peirano G, Pillai DR, Pitondo-Silva A, Richardson D, Pitout JD. The characteristics of NDM-producing Klebsiella pneumoniae from Canada. Diagn Microbiol Infect Dis. 2011;71:106–9. DOIPubMedGoogle Scholar
- Sidjabat H, Nimmo GR, Walsh TR, Binotto E, Htin A, Hayashi Y, Carbapenem resistance in Klebsiella pneumoniae due to the New Delhi metallo-β-lactamase. Clin Infect Dis. 2011;52:481–4. DOIPubMedGoogle Scholar
- Kitchel B, Sundin DR, Patel JB. Regional dissemination of KPC-producing Klebsiella pneumoniae. Antimicrob Agents Chemother. 2009;53:4511–3. DOIPubMedGoogle Scholar
- Coque TM, Novais A, Carattoli A, Poirel L, Pitout J, Peixe L, Dissemination of clonally related Escherichia coli strains expressing extended-spectrum β-lactamase CTX-M-15. Emerg Infect Dis. 2008;14:195–200. DOIPubMedGoogle Scholar
- Nordmann P, Poirel L, Toleman MA, Walsh TR. Does broad-spectrum β-lactam resistance due to NDM-1 herald the end of the antibiotic era for treatment of infections caused by gram-negative bacteria? J Antimicrob Chemother. 2011;66:689–92. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 19, Number 6—June 2013
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
J. Kamile Rasheed, Division of Healthcare Quality Promotion, Centers for Disease Control and Prevention, 1600 Clifton Rd NE, Mailstop G08, Atlanta, GA 30329, USA
Top