Volume 22, Number 11—November 2016
Research
Global Escherichia coli Sequence Type 131 Clade with blaCTX-M-27 Gene
Cite This Article
Citation for Media
Abstract
The Escherichia coli sequence type (ST) 131 C2/H30Rx clade with the blaCTX-M-15 gene had been most responsible for the global dissemination of extended-spectrum β-lactamase (ESBL)–producing E. coli. ST131 C1/H30R with blaCTX-M-27 emerged among ESBL-producing E. coli in Japan during the late 2000s. To investigate the possible expansion of a single clade, we performed whole-genome sequencing for 43 Japan and 10 global ST131 isolates with blaCTX-M-27 (n = 16), blaCTX-M-14 (n = 16), blaCTX-M-15 (n = 13), and others (n = 8). We also included 8 ST131 genomes available in public databases. Core genome-based analysis of 61 isolates showed that ST131 with blaCTX-M-27 from 5 countries formed a distinct cluster within the C1/H30R clade, named C1-M27 clade. Accessory genome analysis identified a unique prophage-like region, supporting C1-M27 as a distinct clade. Our findings indicate that the increase of ESBL-producing E. coli in Japan is due mainly to emergence of the C1-M27 clade.
The global increase in resistance to the third-generation cephalosporins and fluoroquinolones among extraintestinal pathogenic Escherichia coli (ExPEC) is a public health concern because of the importance of these drugs in treating serious infections (1). The extended-spectrum β-lactamases (ESBLs), especially CTX-M types, contribute to third-generation cephalosporin resistance among ExPEC, and specific mutations in quinolone resistance–determining regions in gyrA and parC mainly contribute to fluoroquinolone resistance (2). The increase in resistance among ExPEC has resulted mainly from the recent expansion of a pandemic clonal group known as E. coli sequence type (ST) 131, which is usually multidrug resistant and is associated with CTX-M-15, the most prevalent β-lactamase among ESBL-producing ExPEC (2). ST131 harbors more virulence factors than other antimicrobial-resistant ExPEC and can cause severe infections (2,3).
Recent studies using whole-genome sequencing (WGS) analysis revealed that ST131 comprises different lineages or clades (4,5). Price et al. found a dominant fluoroquinolone-resistant lineage (named H30R) in North America that contains the fimH 30 allele and was associated with characteristic quinolone resistance–determining region mutations (2,4). ST131 with the blaCTX-M-15 gene formed a distinct cluster within the H30R lineage, referred to as the H30Rx clade (4). Petty et al. confirmed these findings using a collection of strains from 6 countries (5). In their study, H30R and H30Rx clades correspond to clade C and clade C2 (subset of clade C), respectively. The other clade C subset, clade C1, included ST131 isolates with different CTX-Ms than blaCTX-M-15.
Globally, the CTX-M-15–producing C2/H30Rx clade is mostly responsible for the pandemic of ExPEC with ESBLs (2), but in Japan, ExPEC with blaCTX-M-15 is rare despite the predominance of ST131 among ESBL-producing isolates (6). Before 2005, ST131 C1/H30R negative for Rx containing blaCTX-M-14 predominated among Japanese ST131 (6). In 2006, ST131 C1/H30R with blaCTX-M-27 was detected in Japan, and the numbers of this lineage escalated since 2010 and are responsible for the substantial increase of ESBL-producing ExPEC in Japan (6). Moreover, blaCTX-M-27 is confined to ST131, whereas other CTX-Ms, such as blaCTX-M-14 and blaCTX-M-15, are equally present among ST131 and non-ST131 E. coli isolates (3).
blaCTX-M-27 is an infrequent global blaCTX-M allele that differs by only 1 nt from blaCTX-M-14, which results in 1 aa change at position 240 (1,6). ST131 with CTX-M-27 had previously been reported from other countries, such as Korea (isolation year 2008), China (2013–2014), Australia (2009–2010), Nepal (2013–2014), Cambodia (2004–2005), Israel (2008–2009), Czech Republic (2008–2011), Switzerland (2011), Spain (2012), France (2012), Portugal (2013–2014), Netherlands (2011), Canada (2005), and United States (2013) (2, 5–15). Because of the rapid increase of CTX-M-27–producing ST131 in Japan since 2010 (6), we designed a study to characterize these isolates using WGS techniques.
Bacterial Isolates
We selected 43 nonduplicate ST131 clinical isolates collected from 2 multicenter surveillance programs in Japan for WGS to represent 3 major ESBL-producing ST131 (CTX-M-27–producing H30R, 13 isolates; CTX-M-14–producing H30R, 9 isolates; CTX-M-15–producing H30Rx, 11 isolates) and other ST131 (CTX-M-14+CTX-M-15–producing H30Rx, 2 isolates; CTX-M-14–producing H30Rx, 1 isolate; CTX-M-14–producing H22, 1 isolate; CTX-M-2–producing H22, 1 isolate; TEM-producing H30, 2 isolates; non–ESBL-producing H30R, 3 isolates) in Japan (6) (Table). One of the surveillance programs collected ESBL-producing E. coli isolates during 2001–2010 at 10 acute-care hospitals in the Kyoto and Shiga prefectures of Japan (6); the other program collected all E. coli isolates during December 2014 at 10 acute-care hospitals in the 5 prefectures in central Japan. ST131 isolates were identified by PCR specific for mdh and gyrB alleles, O25b or O16 rfb variants, fimH allele, and H30Rx status (6). The selection process of the Japanese ST131 ensured equal representation by geographic location, specimen type, and date of isolation.
In addition to isolates from Japan, we obtained 10 CTX-M–producing ST131 isolates from global collections that previously had been characterized by multilocus sequence typing (MLST) (Table; Technical Appendix Table 1). We selected all of the CTX-M-27 producers, 1 CTX-M-14 producer per country, and 2 CTX-M-15 producers. We also sought public databases for ST131 H30 and included sequence data for 8 isolates from countries other than Japan: CTX-M-27 producers (3 raw reads, 2 draft genomes); CTX-M-14 producer (1 raw read); and CTX-M-15–producing C2/H30Rx (2 complete genomes) (Table; Technical Appendix Table 1) (5,9,16–18).
WGS
We used the Nextera XT DNA sample preparation kit (Illumina, San Diego, CA, USA) to prepare libraries for sequencing. Samples were multiplexed and sequenced on an Illumina MiSeq for 600 cycles (300-bp paired-end) or NextSeq500 for 300 cycles (151-bp paired-end). The ST131 genomes were sequenced at an average depth of 44.03 (SD ± 14.70) and an average coverage of 97.73% (SD ± 0.93%) using the 5,109,767-bp EC958 chromosome as previously described (16).
Core Genome Analysis
We used a core genome single-nucleotide polymorphism (SNP)–based approach to create a phylogenetic tree. We identified SNPs using raw read mapping followed by duplicate read removal, realignment, quality score recalibration, and variant filtering (Technical Appendix). Reads from 53 isolates sequenced in this study and 4 isolates (S100EC, S107EC, S108EC, and S135EC) (5) were aligned against a reference genome of EC958, and SNPs were called. The remaining 4 draft or complete genomes underwent whole-genome alignment against EC958 to make EC958-like pseudo-chromosomes that contained only SNPs. The SNP-only core genome was identified as the blocks of >500 bp common to all 61 study isolates to ensure that each block represented a common segment from good alignment in each isolate and that the block had enough length to enable identification (5). A maximum-likelihood tree was built using RAxML (19). A recombination-free tree was also build by excluding recombination sites identified using a Bayesian analysis software BRATNextGen (20).
Comparative Genomic Analysis
To define presence of genes and their alleles, we used SRST2 with trimmed reads or BLAST+ (executables [http://blast.ncbi.nlm.nih.gov/]) with assembled draft genomes and following databases or typing schemes: ResFinder antimicrobial resistance gene database, VFDB and VirulenceFinder virulence gene databases, serotypeFinder O:H typing database, PlasmidFinder plasmid replicon database, MLST (http://mlst.ucc.ie/mlst/dbs/Ecoli), plasmid MLST, fimH typing, gyrA/parC typing, ST131 virotyping, and detection of H30Rx-specific ybbW SNPs, plasmid addiction systems, and blaCTX-M genetic environment (online Technical Appendix). We used pangenome analysis to identify clade specific segments among draft or complete genomes. BRIG was used to visualize similarity of genomes to ST131 genomic islands (16) and to the ST131 reference plasmid pEC958 (21).
Statistical Analysis and Sequence Data Accession Numbers
We compared categorical variables using Fisher exact test. A p value <0.05 was considered statistically significant. We conducted our statistical analysis using Stata, version 13.1 (StataCorp, College Station, TX, USA). The sequences were deposited in the DDBJ Sequence Read Archive database (accession no. DRA004266 and DRA004267).
Bacterial Isolates
The study comprised 60 clinical and 1 environmental ST131 isolates (Table; Technical Appendix Table 1). We confirmed the types of β-lactamase genes, ST131 status, fimH allele numbers, and H30Rx status using draft genomes.
Core Genome SNP-based Phylogenetic Tree
We identified a 4,086,650-bp core genome that included 5,280 SNPs by mapping and alignment of the 61 study isolates to EC958 (Figure 1). The ciprofloxacin-resistant isolates with gyrA 1AB and parC 1aAB alleles formed the C/H30R cluster that comprised the C2/H30Rx and C1/H30R clades. The C2/H30Rx clade included isolates with blaCTX-M-15 (n = 15) and blaCTX-M-14 (n = 1) and isolates with both blaCTX-M-15 and blaCTX-M-14 (n = 2) (Figure 1). The C1/H30R clade included isolates with blaCTX-M-27 (n = 21) and blaCTX-M-14 (n = 14) and isolates without ESBLs (n = 3) (Figure 1). Within the C1/H30R clade, 19 of 21 CTX-M-27–producing isolates clustered into a distinct group, named the C1-M27 clade (Figure 1). E. coli ST131 C1-M27 comprised isolates from Japan (n = 13; isolation years 2004–2014), Australia (n = 2; 2009–2010), United States (n = 2; 2013–2014), Canada (n = 1, 2008), and Thailand (n = 1, 2013).
Analysis of the core genome showed that 79 segments (i.e., 304,782 bp, including 3,453 SNPs) were associated with recombination sites (Technical Appendix Figure 1). This finding suggests that recombinant segments contained 65% of SNPs with subsequent higher frequency of SNPs compared with nonrecombinant regions (average 11 vs. 0.48 SNPs/kb, respectively). The phylogenetic tree created without recombination sites showed the same results as the phylogenetic tree obtained with recombination sites (Technical Appendix Figure 2). In addition to the core genome–based phylogeny with or without recombination sites, the C1-M27 clade was defined by a unique accessory genome of the M27PP1.
The C1-M27 Clade–Specific Region
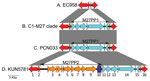
The pangenome analysis of genomes from all the isolates identified an 11,894-bp region named M27PP1 that was specific to all the isolates from the C1-M27 clade. Further analysis using the BLAST database and Sanger sequencing for gap filling showed that this region was identical to a prophage-like genomic island (GenBank accession no. CP006632) in E. coli PCN033 that belonged to phylogenetic group D and was isolated from a pig in China. The BLAST database also identified 2 similar sequences (i.e., 99.9% homology): A CMY-2 containing plasmid pEQ011 (GenBank accession no. NC_023315) in an E. coli isolate from a horse in Ireland (22) and a multidrug-resistant plasmid pSD853_88 (GenBank accession no. JF267652) found in a bovine Salmonella enterica isolate in the United States. M27PP1 was inserted into chromosome creating a 7-bp direct repeat region (Figure 2). PCN033 had the same flanking structure as the M27PP1, whereas the 2 plasmids (pEQ011, pSD853_89) contained only a 44-bp similar segment at 5′ side and other parts of these plasmids were not found in the C1-M27 clade isolates.
Two E. coli ST131 C1-M27 isolates (i.e., KUN5781 and Ec 24) had an additional insertion region of 19,352 bp, named M27PP2, situated upstream of M27PP1. M27PP2 was accommodated within the same 7-bp direct repeat region (Figure 2). M27PP2 included a 15,555-bp region that showed 88.9% homology to a prophage-like sequence in the chromosome of the γ proteobacterium HdN1 (GenBank accession no. FP929140) and 99.8% homology to the insertion element ISSen4.
Genomic Comparison of the ST131 Genomic Islands and Virulence Genes
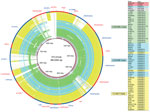
The sequences of the study isolates were similar to the ST131 genomic islands in EC958 and JJ1886 (a CTX-M-15–producing C2/H30Rx strain obtained in the United States from a patient with fatal urosepsis) (Figure 3) (17). The C1-M27 clade isolates lacked the prophage 1 region present in EC958 (Figure 3). This prophage 1 region, specific for ST131, was present among the non–C1-M27 ST131 isolates in this study, except for BRG23 and EcSA01. The presence of ExPEC-associated virulence genes is shown in Technical Appendix Figure 3. The senB enterotoxin gene was more common in C1/H30R (than in C2/H30Rx). No significant differences existed in the distribution of virulence genes between E. coli ST131 C1-M27 and other isolates.
Plasmid Replicons, Addiction Systems, and Antimicrobial Drug Resistance Genes
We compared the study isolates with pEC958, the plasmid present in EC958 that carries blaCTX-M-15 (Technical Appendix Figure 4). The C1-M27 clade lacked the first part of the transfer region (tra) present in pEC958. Some regions common to both C2/H30Rx and C1/H30R clades were present in pEC958. The C1/H30R clade producing CTX-M-27 or CTX-M-14 (including C1-M27) contained mostly F1:A2:B20 replicons, whereas the C2/H30Rx clade producing CTX-M-15 contained mainly F2:A1:B- replicons (Technical Appendix Figure 5). The C1-M27 clade was negative for Tn2 containing blaTEM-1. Two C1-M27 isolates from Thailand and the United States were also positive for blaNDM-1 (Technical Appendix Figure 5).
A previously unreported clade named C1-M27 within C1/H30R clade is responsible for the epidemic of ESBL-producing ExPEC in Japan and has already been disseminated to 5 countries on 3 continents. ST131 containing blaCTX-M-27 responsible for human infections has been reported from various continents (2) and is especially common among ESBL-producing ExPEC in certain countries, such as Israel, the Czech Republic, and Switzerland (2,13,14). CTX-M-27–producing ST131 also is present among nonclinical and nonhuman E. coli isolates, including fecal specimens of healthy children attending day care centers in France; fecal specimens of healthy adults in China, Portugal, and the Netherlands; samples from sick dogs and cats in Japan; samples from water birds from central Europe and Swiss rivers and lakes; and samples of well water from China (2,10,11,15,23–25). The most common ESBL among E. coli ST131 in nonhuman samples is CTX-M-27 (2,23–25). ST131 with blaCTXM-15 is rare among animal and environmental E. coli isolates (26). Our analysis of IEH71520, an E. coli isolate from vacuum cleaner dust in the United States (15), showed that this ST131 isolate belong to the C1-M27 clade. The C1-M27 clade is likely to be present among animal and environmental ST131, and such isolates might act as a hidden reservoir for the introduction of ST131 containing blaCTX-M-27 into human medicine.
E. coli ST131 C1-M27 had an additional, unique prophage-like region (M27PP1) within its chromosome, lacked the prophage 1 genomic island previously identified in ST131 C2/H30Rx, and were negative for the transposon Tn2 containing blaTEM-1 (Figure 3; Technical Appendix Figure 5). The direct flanking repeat sequences surrounding M27PP1 suggest that this region was introduced into E. coli ST131 C1/H30R with blaCTX-M-27 by a recombination event that was then followed by the clonal expansion of the C1-M27 clade.
Recent studies focusing on evolutionary history of ST131 suggested that C1/H30R and C2/H30Rx clades emerged ≈30 years ago, after their acquisition of gyrA-1AB and parC-1aAB alleles from C0/H30 (non-R) clade (27,28). The phylogeny and smaller numbers of SNPs in the C1-M27 clade (Figure 1) suggest that this clade was recently diverged from the C1/H30R. In the time-scaled phylogeny presented by Stoesser et al. (27), a cluster that included 6 CTX-M-27–producing isolates from Cambodia, Thailand, and Laos in 2007–2011 was present within the C1/H30R clade. This cluster, supposed to be the C1-M27 clade, diverged in the early 2000s, supporting our hypothesis.
CTX-M-27–producing ST131 that belongs to the H41 lineage previously had been described from Japan (6) and China (15). The characterization of the Japanese ST131 H41 isolates showed different genetic structures flanking the blaCTX-M-27 from those structures present in E. coli ST131 H30R (6). The flanking regions previously characterized in ST131 H41 were identical to the flanking regions in ST131 non–C1-M27 from this study. It seems there are 2 types of structures flanking the blaCTX-M-27 among E. coli ST131; 1 type is confined to clade C1-M27 (i.e., 208 bp of ΔISEcp1 upstream and ΔIS903D downstream), whereas another structure (i.e., 388 bp of ΔISEcp1 upstream and full IS903D downstream) is distributed among non–C1-M27 isolates, including ST131 H41 (6). Therefore, ST131 H41, through horizontal transfer of blaCTX-M-27, is unlikely to have played a substantial role in the emergence of the C1-M27 clade.
Two ST131 isolates with blaCTX-M-27 from Australia and Vietnam did not belong to the C1-M27 clade (Figure 1). These 2 isolates differ from the C1-M27 clade in that their core genomes had more SNPs (158 vs. 68), contained the prophage 1 ST131-specific region, and lacked the M27PP1 and M27PP2 elements. Moreover, the genetic environment surrounding the blaCTX-M-27 differed from E. coli ST131 C1-M27 (as described previously). The isolate from Vietnam lacked mph(A)-mrx-mphR, tetR-tet(A), sul2-strA-strB, and In54 resistance genes, compared with the C1-M27 clade (Technical Appendix Figure 5). These differences indicate that some ST131 isolates might acquire blaCTX-M-27 independently from the C1-M27 clade.
Our study has several limitations. Most isolates originated from Japan. However, we were able to include ST131 C1-M27 isolates from 5 countries on 3 continents and C1/H30R isolates producing CTX-M-14 or CTX-M-15 from 6 countries on 4 continents. Another limitation was that we were able to obtain only 1environmental ST131 isolate with blaCTX-M-27 (IEH71520). Future studies that include environmental isolates will provide additional insights into molecular epidemiology and evolutionary history of the C1-M27 clade. We could not analyze plasmid contents of blaCTX-M-27 because blaCTX-M-27–containing contigs were too short. The sequencing of plasmids that contain blaCTX-M-27 obtained from various ST131 clades (including the C1-M27 clade) should also be undertaken.
In conclusion, we showed that the recent increase in ESBL-producing E. coli from Japan resulted from emergence of a ST131 C1/H30R subclade with blaCTX-M-27. This clade, named C1-M27, had unique genomic characteristics and was present in ST131 from Thailand, Australia, Canada, and the United States. Our findings suggest that the C1-M27 clade is contributing to the global success of ST131. E. coli ST131 C1-M27 poses a major new public health threat because of its global distribution and association with the very dominant C/H30 lineage. We urgently need rapid cost-effective detection methods for E. coli ST131 C1-M27 and well-designed epidemiologic and molecular studies to understand the dynamics of transmission, risk factors, and reservoirs for ST131 C1-M27. These efforts will provide insight into the emergence and spread of this multidrug-resistant clade that will lead to information essential for preventing the spread of ST131.
Dr. Matsumura is an assistant professor at the Clinical Laboratory Medicine, Kyoto University Graduate School of Medicine, Kyoto, Japan. His main research focuses on the detection and molecular epidemiology of antimicrobial drug resistance mechanisms among gram-negative bacteria.
Acknowledgments
We thank the members of the Kyoto-Shiga Clinical Microbiology Study Group (Naohisa Fujita, Toshiaki Komori, Yukiji Yamada, Tsunehiro Shimizu, Akihiko Hayashi, Tamotsu Ono, Harumi Watanabe, Naoko Fujihara, Takeshi Higuchi, Kunihiko Moro, Masayo Shigeta, Kaneyuki Kida, Hiromi Terada, Fusayuki Tsuboi, Yoshihisa Sugimoto, and Chiyoko Fukumoto) and the members of the 89JAID–BRG (89th Annual Meeting of the Japanese Association for Infectious Diseases–drug-resistant Bacteria Research Group; Ooi Yukimasa, Tomohiro Higashiyama, Masaaki Sasano, Harumi Watanabe, Tsunehiro Shimizu, Akihiko Hayashi, Yasufumi Matsumura, Taro Noguchi, Miki Nagao, Masaki Yamamoto, Michio Tanaka, Takeshi Higuchi, Toru Kanahashi, Naohisa Fujita, Toshiaki Komori, Yukiji Yamada, Kohei Ueda, Go Yamamoto, Kunihiko Moro, and Shigenobu Yabu) for collection of isolates. We also thank Sayo Shitashiro and Karen Poon for technical assistance.
This work was supported by Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Young Scientists (B) Grant No. 15K19179 (Y.M.), the John Mung Program from Kyoto University, Japan (Y.M.), and a research grant from the Calgary Laboratory Services (#10006465; J.D.D.P). J.D.D.P had previously received research funds from Merck and Astra Zeneca.
References
- Pitout JD, Laupland KB. Extended-spectrum beta-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet Infect Dis. 2008;8:159–66.DOIPubMedGoogle Scholar
- Nicolas-Chanoine MH, Bertrand X, Madec JY. Escherichia coli ST131, an intriguing clonal group. Clin Microbiol Rev. 2014;27:543–74.DOIPubMedGoogle Scholar
- Matsumura Y, Yamamoto M, Nagao M, Ito Y, Takakura S, Ichiyama S; Kyoto-Shiga Clinical Microbiology Study Group. Association of fluoroquinolone resistance, virulence genes, and IncF plasmids with extended-spectrum-β-lactamase-producing Escherichia coli sequence type 131 (ST131) and ST405 clonal groups. Antimicrob Agents Chemother. 2013;57:4736–42.DOIPubMedGoogle Scholar
- Price LB, Johnson JR, Aziz M, Clabots C, Johnston B, Tchesnokova V, The epidemic of extended-spectrum-β-lactamase-producing Escherichia coli ST131 is driven by a single highly pathogenic subclone, H30-Rx. MBio. 2013;4:e00377–13. DOIPubMedGoogle Scholar
- Petty NK, Ben Zakour NL, Stanton-Cook M, Skippington E, Totsika M, Forde BM, Global dissemination of a multidrug resistant Escherichia coli clone. Proc Natl Acad Sci U S A. 2014;111:5694–9.DOIPubMedGoogle Scholar
- Matsumura Y, Johnson JR, Yamamoto M, Nagao M, Tanaka M, Takakura S, ; Kyoto–Shiga Clinical Microbiology Study Group; Kyoto-Shiga Clinical Microbiology Study Group. CTX-M-27- and CTX-M-14-producing, ciprofloxacin-resistant Escherichia coli of the H30 subclonal group within ST131 drive a Japanese regional ESBL epidemic. J Antimicrob Chemother. 2015;70:1639–49.PubMedGoogle Scholar
- Sherchan JB, Hayakawa K, Miyoshi-Akiyama T, Ohmagari N, Kirikae T, Nagamatsu M, Clinical epidemiology and molecular analysis of extended-spectrum-β-lactamase-producing Escherichia coli in Nepal: characteristics of sequence types 131 and 648. Antimicrob Agents Chemother. 2015;59:3424–32.DOIPubMedGoogle Scholar
- Clermont O, Dhanji H, Upton M, Gibreel T, Fox A, Boyd D, Rapid detection of the O25b-ST131 clone of Escherichia coli encompassing the CTX-M-15-producing strains. J Antimicrob Chemother. 2009;64:274–7.DOIPubMedGoogle Scholar
- McGann P, Snesrud E, Ong AC, Appalla L, Koren M, Kwak YI, War wound treatment complications due to transfer of an IncN plasmid harboring bla(OXA-181) from Morganella morganii to CTX-M-27-producing sequence type 131 Escherichia coli. Antimicrob Agents Chemother. 2015;59:3556–62.DOIPubMedGoogle Scholar
- Rodrigues C, Machado E, Fernandes S, Peixe L, Novais Â. An update on faecal carriage of ESBL-producing Enterobacteriaceae by Portuguese healthy humans: detection of the H30 subclone of B2-ST131 Escherichia coli producing CTX-M-27. J Antimicrob Chemother. 2016;71:1120–2.DOIPubMedGoogle Scholar
- Reuland EA, Al Naiemi N, Kaiser AM, Heck M, Kluytmans JA, Savelkoul PH, Prevalence and risk factors for carriage of ESBL-producing Enterobacteriaceae in Amsterdam. J Antimicrob Chemother. 2016;71:1076–82.DOIPubMedGoogle Scholar
- Peirano G, van der Bij AK, Freeman JL, Poirel L, Nordmann P, Costello M, Characteristics of Escherichia coli sequence type 131 isolates that produce extended-spectrum β-lactamases: global distribution of the H30-Rx sublineage. Antimicrob Agents Chemother. 2014;58:3762–7.DOIPubMedGoogle Scholar
- Micenková L, Sišková P, Bosák J, Jamborová I, Cernohorská L, Smajs D. Characterization of human uropathogenic ESBL-producing Escherichia coli in the Czech Republic: spread of CTX-M-27-producing strains in a university hospital. Microb Drug Resist. 2014;20:610–7.DOIPubMedGoogle Scholar
- Adler A, Gniadkowski M, Baraniak A, Izdebski R, Fiett J, Hryniewicz W, ; MOSAR WP5 and WP2 study groups. Transmission dynamics of ESBL-producing Escherichia coli clones in rehabilitation wards at a tertiary care centre. Clin Microbiol Infect. 2012;18:E497–505.DOIPubMedGoogle Scholar
- Zhong YM, Liu WE, Liang XH, Li YM, Jian ZJ, Hawkey PM. Emergence and spread of O16-ST131 and O25b-ST131 clones among faecal CTX-M-producing Escherichia coli in healthy individuals in Hunan Province, China. J Antimicrob Chemother. 2015;70:2223–7.DOIPubMedGoogle Scholar
- Forde BM, Ben Zakour NL, Stanton-Cook M, Phan MD, Totsika M, Peters KM, The complete genome sequence of Escherichia coli EC958: a high quality reference sequence for the globally disseminated multidrug resistant E. coli O25b:H4-ST131 clone. PLoS One. 2014;9:e104400.DOIPubMedGoogle Scholar
- Andersen PS, Stegger M, Aziz M, Contente-Cuomo T, Gibbons HS, Keim P, Complete genome sequence of the epidemic and highly virulent CTX-M-15–producing H30-Rx subclone of Escherichia coli ST131. Genome Announc. 2013;1:e00988–13.DOIPubMedGoogle Scholar
- Kutumbaka KK, Han S, Mategko J, Nadala C, Buser GL, Cassidy MP, Draft denome sequence of blaNDM-1-positive Escherichia coli O25b-ST131 clone isolated from an environmental sample. Genome Announc. 2014;2:e00462–14.DOIPubMedGoogle Scholar
- Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3.DOIPubMedGoogle Scholar
- Marttinen P, Hanage WP, Croucher NJ, Connor TR, Harris SR, Bentley SD, Detection of recombination events in bacterial genomes from large population samples. Nucleic Acids Res. 2012;40:e6.DOIPubMedGoogle Scholar
- Phan MD, Forde BM, Peters KM, Sarkar S, Hancock S, Stanton-Cook M, Molecular characterization of a multidrug resistance IncF plasmid from the globally disseminated Escherichia coli ST131 clone. PLoS One. 2015;10:e0122369.DOIPubMedGoogle Scholar
- Karczmarczyk M, Wang J, Leonard N, Fanning S. Complete nucleotide sequence of a conjugative IncF plasmid from an Escherichia coli isolate of equine origin containing blaCMY-2 within a novel genetic context. FEMS Microbiol Lett. 2014;352:123–7.DOIPubMedGoogle Scholar
- Zhang H, Zhou Y, Guo S, Chang W. Prevalence and characteristics of extended-spectrum beta-lactamase (ESBL)-producing Enterobacteriaceae isolated from rural well water in Taian, China, 2014. Environ Sci Pollut Res Int. 2015;22:11488–92.DOIPubMedGoogle Scholar
- Müller A, Stephan R, Nüesch-Inderbinen M. Distribution of virulence factors in ESBL-producing Escherichia coli isolated from the environment, livestock, food and humans. Sci Total Environ. 2016;541:667–72.DOIPubMedGoogle Scholar
- Harada K, Nakai Y, Kataoka Y. Mechanisms of resistance to cephalosporin and emergence of O25b-ST131 clone harboring CTX-M-27 β-lactamase in extraintestinal pathogenic Escherichia coli from dogs and cats in Japan. Microbiol Immunol. 2012;56:480–5.DOIPubMedGoogle Scholar
- Mathers AJ, Peirano G, Pitout JD. The role of epidemic resistance plasmids and international high-risk clones in the spread of multidrug-resistant Enterobacteriaceae. Clin Microbiol Rev. 2015;28:565–91.DOIPubMedGoogle Scholar
- Stoesser N, Sheppard AE, Pankhurst L, De Maio N, Moore CE, Sebra R, ; Modernizing Medical Microbiology Informatics Group (MMMIG). Evolutionary history of the global emergence of the Escherichia coli epidemic clone ST131. MBio. 2016;7:e02162–15. DOIPubMedGoogle Scholar
- Ben Zakour NL, Alsheikh-Hussain AS, Ashcroft MM, Khanh Nhu NT, Roberts LW, Stanton-Cook M, Sequential acquisition of virulence and fluoroquinolone resistance has shaped the evolution of Escherichia coli ST131. MBio. 2016;7:e00347–16.PubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 22, Number 11—November 2016
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Yasufumi Matsumura, Department of Clinical Laboratory Medicine, Kyoto University Graduate School of Medicine, 54 Shogoinkawahara-cho, Sakyo-ku, Kyoto 6068507, Japan
Top