Volume 24, Number 12—December 2018
Research
Survey of Ebola Viruses in Frugivorous and Insectivorous Bats in Guinea, Cameroon, and the Democratic Republic of the Congo, 2015–2017
Cite This Article
Citation for Media
Abstract
To clarify the role of bats in the ecology of Ebola viruses, we assessed the prevalence of Ebola virus antibodies in a large-scale sample of bats collected during 2015–2017 from countries in Africa that have had previous Ebola outbreaks (Guinea, the Democratic Republic of the Congo) or are at high risk for outbreaks (Cameroon). We analyzed 4,022 blood samples of bats from >12 frugivorous and 27 insectivorous species; 2–37 (0.05%–0.92%) bats were seropositive for Zaire and 0–30 (0%–0.75%) bats for Sudan Ebola viruses. We observed Ebola virus antibodies in 1 insectivorous bat genus and 6 frugivorous bat species. Certain bat species widespread across Africa had serologic evidence of Zaire and Sudan Ebola viruses. No viral RNA was detected in the subset of samples tested (n = 665). Ongoing surveillance of bats and other potential animal reservoirs are required to predict and prepare for future outbreaks.
Since the first outbreak of Ebola virus disease (EVD) in 1976 in the northern part of the Democratic Republic of the Congo (DRC), 26 recognized outbreaks have occurred in humans across Africa; fatality rates of outbreaks have been 25%–90% (1–4). Each EVD outbreak most likely resulted from independent zoonotic events.
Bats are believed to play a role in the ecology of Ebola viruses as a reservoir species (5). Bats might infect humans directly or via intermediate amplifying hosts, like nonhuman primates or duikers (6,7). Bats might serve as a source of infection in certain areas where bats are hunted and eaten as bushmeat, but infection could also occur after consumption of fruits contaminated with saliva, urine, or feces from Ebola virus–infected bats (8,9). Ebola virus emergence through exposure to bats was suspected for at least 2 outbreaks: Luebo (the DRC) in 2007 and West Africa in 2013 (10,11).
Relatively few data are available to support the role of bats in the ecology of Ebola viruses. During the EVD outbreaks of 2003 in Gabon and the Congo, Zaire Ebola virus RNA and antibodies were detected in live-caught specimens from 3 fruit bat species (Epomops franqueti, Hypsignathus monstrosus, Myonycteris torquata); virus sequences were found in the livers or spleens of a few bats (6). In subsequent studies in Gabon, the Congo, Ghana, and Zambia, antibodies were detected in additional frugivorous bat species (Eidolon helvum, Epomophorus gambianus, Rousettus aegyptiacus, Micropteropus pusillus) and 1 insectivorous species (Mops condylurus) (12–16). The amplification and sequencing of viral RNA of other filoviruses in bats, such as Marburg virus in bats from Africa (17–20), Lloviu virus in bats from Europe (21), and new filoviruses in bats from China (22), has provided additional evidence for a possible role of bats in Ebola virus ecology.
In general, EVD outbreaks have been limited in terms of their geographic spread and chains of human-to-human transmission (1). However, during the 2013–2016 outbreak, virus spread to the urban areas of 3 countries, infecting ≈30,000 persons in Guinea, Sierra Leone, and Liberia, and ≈11,000 deaths were recorded (23). This outbreak illustrated the potential for epidemic spread from a single zoonotic transmission, with severe public health and socioeconomic impact (24). Additional studies are urgently needed to identify the animal reservoir, predict EVD outbreak risks, and improve our capacity to control epidemics.
In previous modeling studies, areas were defined as at risk for EVD outbreaks on the basis of data collected from a limited number of wildlife bat species from a few geographic regions (5,25). Also, a wide variety of serologic assays and interpretation criteria have been used, making comparison of results challenging (12–16,26,27). For this study, we performed a large serosurvey with a highly specific and sensitive high-throughput assay to assess Ebola virus prevalence in bats from Africa (28). We studied bats from Guinea and the DRC, countries with previous EVD outbreaks, and Cameroon, a country considered at high risk for future EVD outbreaks (5,25).
Study Sites and Sample Collection
During November 2015–August 2017, we collected samples from free-ranging frugivorous and insectivorous bats in Guinea, Cameroon, and the DRC. We captured bats at night using ground mist nets or harp traps in roosting and foraging sites. We set up ground mist nets (12 × 3.2 m) of 30-mm and 60-mm mesh sizes at different heights (1–7 m) to maximize capture of different species. We opened nets or harp traps just before sunset and checked for bats every 1–2 hours. Captured bats were released the same night immediately after sampling. Using bat whole blood taken by venipuncture of the propatagial or brachial vein, we dropped blood samples directly onto Whatman 903 filter paper (GE Healthcare, Feasterville-Trevose, PA, USA). We air-dried and preserved samples individually in plastic bags containing silica desiccant and stored them in hermetic boxes; 2–3 weeks later, we transferred dried blood spots to −20°C until needed for analysis. Data recorded in the field included information on capture site (global positioning system coordinates, ecologic environment), capture method, morphology (body measurements, weight, color), sex, age class (adult, juvenile), and species (identified visually). We collected negative control samples (n = 145) from a captive-born insectivorous bat species (103 Carollia perspicillata bats) hosted at the Parc Zoologique de Montpellier (Montpellier, France) and 2 frugivorous bat species (19 Pteropus giganteus bats, 23 R. aegyptiacus bats) hosted at Wilhelma Zoo and Botanical Garden (Stuttgart, Germany). We collected and preserved samples the same way we did for free-ranging bats.
Screening for Ebola Virus Antibodies
We tested dried blood spots with a Luminex-based serologic assay adapted for bats (28) (Technical Appendix). The assay included recombinant Ebola virus proteins glycoprotein, nucleoprotein, or viral protein 40 for different lineages: Zaire, Sudan, Bundibugyo, and Reston. We reconstituted plasma from dried blood spots as previously described (28) and incubated 100 μL of sample (final plasma dilution 1:2,000) with 50 µL of recombinant protein–coated beads (2 µg protein/1.25 × 106 beads) in 96-well flat-bottom filter plates (Millipore, Tullagreen, Ireland) on a plate shaker at 300 rpm for 16 h at 4°C in the dark. After washing, we added 0.1 μg/mL of goat anti–bat biotin–labeled IgG (Euromedex, Souffelweyersheim, France) per well and incubated for 30 min at 300 rpm. After another round of washing, we added 50 µL of 4 µg/mL streptavidin-R-phycoerythrin (Fisher Scientific, Illkirch, France) per well and incubated for 10 min at 300 rpm. Reactions were read with BioPlex-200 (BioRad, Marnes-la-Coquette, France). We expressed results as median fluorescence intensity (MFI) per 100 beads. We included 3 samples on every plate to validate interassay repeatability.
Determination of Cutoffs
In the absence of positive control samples, we used 4 different statistical methods to determine the MFI cutoff value for each antigen (29,30) (Technical Appendix Table 1). First, we used a general formula that involved the MFI of the 145 negative control samples, and we assigned the cutoff as mean plus 4 times the SD (mean + 4×SD). Second, we used a change point analysis (31) to identify the value at which statistical properties of the underlying probability distribution changed. This value was used to identify outliers and classify them as reactive. We used the R package changepoint (32) to calculate a single shift in the arithmetic mean with the at-most-1-change method (33). Third, we fitted univariate distributions to our data and defined the cutoff as a 0.001 risk for error, as was used in other virus serology studies (13,34). We reduced the set of candidate distributions following a bootstrapped skewness-kurtosis analysis (35). We performed fitting by maximum-likelihood estimation and selected the best-fit distribution on the basis of the Akaike information criteria with the R library fitdistrplus (36). A negative binomial distribution best-fit the data; however, we also used the negative exponential distribution as in Pourrut et al. and Laing et al. (13,34). For every antigen, we computed bootstrap values using 10,000 replicates and averaged. We performed analyses with R version 3.3.2 software (https://www.r-project.org/). We considered a blood sample reactive if the MFI of the reaction was above the cutoff. We defined Ebola virus antibody positivity as reactivity to glycoprotein and nucleoprotein of the same lineage, as was done in our previous study (28).
Nucleic Acid Extraction and PCR Screening for Ebola Virus RNA
We extracted total DNA and RNA from dried blood spots as previously described using Nuclisens (bioMerieux, Marcy-l’Etoile, France) or m2000sp methods (Abbott Molecular Inc., Des Plaines, IL, USA), which are known for a high performance recovering nucleic acids from dried blood spots (37,38). For bat species from Cameroon and Guinea, we screened for Zaire Ebola virus RNA by seminested reverse transcription PCR (RT-PCR) targeting the nucleoprotein region of the virus genome. We amplified a 126-bp fragment of Zaire Ebola virus using primers NP1F1 (forward, 5′-CGGACACACAAAAAGAAWGAA-3′) and NP1R-ZR (reverse, 5′-CTCTATCTTKGTGATRTGGCTCTGA-3′) in the first round of PCR and NP1F2 (forward, 5′- TTGTGTGCGARTAACTAYGAGGAAG-3′) plus NP1R-ZR in the second round. For species from the DRC, we performed seminested RT-PCR targeting the viral protein 35 region of the genome using the protocol of He et al. with modifications (41). In the first round, we amplified a 217-bp fragment with primers VP35-F (5′-ATYATGTATGATCACYTVCCWGG-3′) and VP35-R (5′-AGCGRATGTGGATSACRGGT-3′) and, in the second round, a 184-bp product with primers VP35-R and VP35-in-F (5′-GCTTTYCAYCAAYTAGTRCAAG-3′).
Molecular Confirmation of Bat Species
We confirmed bat species identification recorded in the field on a subset of samples by using molecular tests. We amplified an ≈800-bp fragment of mitochondrial cytochrome b using primers cytb-L14724 (forward) and cytb-H15506 (reverse) (11,39,40). We substituted the cytb-L14724 primer with cytb-L140217 (5′-ATGACCAACATCCGAAAATCNCAC-3′) to improve PCR performance for certain species. We purified PCR products through agarose gel (1%) and directly sequenced on an ABI 3500 sequencer (Applied Biosystems, Courtaboeuf, France). We performed BLAST analyses (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to identify the most similar bat species. For samples with no or low similarity (<97%) hits with species in GenBank, we performed phylogenetic analyses with newly obtained sequences and reference sequences for different bat species using maximum-likelihood methods implemented with PhyML (http://www.atgc-montpellier.fr/phyml/) to determine genus.
Bat Species and Sampling
We analyzed blood samples from 4,022 wild bats from 21 different regions in Cameroon (n = 10), Guinea (n = 8), and the DRC (n = 3) (Figure 1; Table 1). To increase species diversity, we captured bats in multiple ecologic settings: forests (49%), open fields (10%), villages (29%), plantations (7%), and urban areas (5%). For 1,470 (36.5%) samples, species identification in the field was confirmed by sequence analysis. At each site, >1 sample was confirmed per sampling date, capture method, and morphologic description. For the remaining samples, species identification was extrapolated by combining molecular and morphologic data, including photographs whenever available. For some insectivorous bat families (Miniopteridae, Molossidae, Nycteridae, Rhinolophidae), identification was possible only at the genus level; for some Molossidae bats, we could not distinguish between Mops and Chaerephon genera because of the lack of sequences in GenBank (Table 2). For 87 (2.16%) samples, species identification was not possible because incomplete data were recorded in the field, and available biologic materials were insufficient for molecular confirmation. We collected samples from 1,736 (43.2%) frugivorous bats (family Pteropodidae) of 12 species and 2,199 (54.7%) insectivorous bats (7 families) of >27 species. The insectivorous bat families sampled, in order of decreasing frequency, were Hipposideridae (31.9%), Molossidae (13.4%), Miniopteridae (5.8%), Rhinolophidae (2.1%), Vespertilionidae (0.8%), Nycteridae (0.5%), and Emballonuridae (0.12%). Overall, 54.7% of bats were female and 43.8% were male; for 1.5% (n = 60) of bats, sex was unknown. Most (77.9%) bats were adults, and 9.6% were juveniles; for 12.5% (n = 502) of bats, age could not be determined or was not recorded.
Bats Antibodies against Different Ebola Virus Antigens
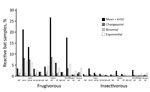
We tested all samples for Ebola virus antibodies. The number of samples reacting with >1 antigen was 734 (18.2%) by the mean + 4×SD method, 274 (6.8%) for the change-point method, 175 (4.4%) for the binomial method, and 457 (11.4%) for the exponential method. Blood samples frequently reacted with glycoprotein antigens; samples reacted most with Zaire and Sudan Ebola virus antigens and least with Reston (Table 3). Simultaneous reactivity to >1 antigen (i.e., glycoprotein, nucleoprotein, viral protein 40) from the same virus lineage was rare. Simultaneous reactivity to the same antigen from different virus lineages was frequent; 32.3%–76.7% of blood samples were reactive to glycoprotein from >2 Ebola virus species, 18.4%–34.0% to viral protein 40, and 1.5%–4.4% to nucleoprotein (Technical Appendix Table 2). When using the criterion simultaneous presence of antibodies to nucleoprotein and glycoprotein, the antibody positivity for Zaire or Sudan Ebola virus antibodies was generally <1% for all bats tested, regardless of cutoff method, and was lower among insectivorous than frugivorous bats: 0.05%–0.27% (insectivorous) and 0.06%–1.79% (frugivorous) for Zaire Ebola virus versus 0%–0.09% (insectivorous) and 0%−1.61% (frugivorous) for Sudan Ebola virus (Table 3; Figure 2). Three samples were positive for Zaire and Sudan Ebola viruses, but only by less stringent cutoff methods (i.e., mean + 4×SD).
Zaire and Sudan Ebola Virus Reactivity of Different Bat Species
We estimated specific reactivity to Zaire and Sudan Ebola viruses by bat species. We did not include Bundibugyo and Reston because recombinant nucleoproteins were not available. Among insectivorous bats, only blood samples from Mops sp. bats (1–6/494) were positive for Zaire or Sudan Ebola virus antibodies (Table 4). Among frugivorous bats, samples from E. helvum, H. monstrosus, and R. aegyptiacus bats had the highest reactivity. We observed Zaire and Sudan Ebola virus seropositivity in these 3 species with almost all cutoff methods: 0.2%–3.3% for Zaire Ebola virus and 1.0%–2.9% for Sudan Ebola virus in E. helvum bat samples, 0.5%–1.6% for Zaire Ebola virus and 1.1%–4.3% for Sudan Ebola virus in H. monstrosus bat samples, and 0.6%–2.5% for Zaire Ebola virus and 0.8%–1.4% for Sudan Ebola virus in R. aegyptiacus bat samples. We observed 2.4% Zaire Ebola virus–seropositive samples for Lissonycteris angolensis bats and 0.5% for Epomophorus sp. bats, but only by less stringent cutoff methods. One sample from M. pusillus bats was seropositive for Sudan Ebola virus. No samples from E. franqueti or M. torquata bats were reactive with any Ebola virus antigens. Samples from the 1 Scotonycteris zenkeri bat and 20 Megaloglossus woermanni bats were seronegative. Overall, Zaire or Sudan Ebola virus antibodies were observed in 7 (1 insectivorous and 6 frugivorous) bat species.
Comparison of Zaire Ebola Virus Seroprevalence in Bats from Africa across Studies
For comparison, we compiled data regarding Zaire Ebola virus serology in bats of known species from previous studies (n = 4,493) and this study (n = 3,935; 46.7%) (Tables 5, 6). Data were available for 3,023 insectivorous bats of ≈30 species from 7 different families; 2,199 (72.7%) were from this study (Table 5). Insectivorous bat samples originated from Guinea, Cameroon, the DRC, and Gabon. Zaire Ebola virus reactivity has been observed only in M. condylurus bat samples from Gabon and Mops sp. bat samples from Cameroon. Data were available for 5,405 frugivorous bats of 17 species from 12 genera from West (Guinea, Ghana), West Central (Cameroon, Gabon, the Congo, the DRC), and East (Zambia) Africa (Table 6). No Zaire Ebola virus reactivity has been seen in blood samples from bat species Casinycteris, Megaloglossus, Nanonycteris, and Scotonycteris, but only a limited number of samples (n = 152) have been tested. Overall, blood samples from 8 frugivorous bat species have been found reactive with Zaire Ebola virus antigens. Blood samples from E. helvum, H. monstrosus, and R. aegyptiacus bats from several countries across Africa have been reported to be seropositive. Reactivity has been observed with samples from E. gambianus bats in Ghana (10.8%) and Guinea. Reactivity was observed with large sample sets from E. franqueti bats derived from Gabon and the Congo and a small sample set from Ghana but not Guinea, Cameroon, or the DRC. M. pusillus and M. torquata bats tested positive for Zaire Ebola virus antibodies in studies in which large sample sets were collected. Among L. angolensis bat samples, only those from Cameroon have tested positive for antibodies.
RT-PCR Screening for Zaire Ebola Virus RNA
We screened 665 samples from the DRC (n = 193), Cameroon (n = 399), and Guinea (n = 73) by RT-PCR for the presence of Zaire Ebola virus RNA. Of the 294 samples originating from bats previously documented to carry Zaire Ebola virus RNA (6) (i.e., H. monstrosus [132 from Cameroon, 1 from the DRC], M. torquata [20 from Cameroon, 25 from the DRC], and E. franqueti [116 from Cameroon]), all were negative for Zaire Ebola virus RNA. Of the 371 samples from bat species E. helvum (58 from Cameroon, 165 from the DRC, 3 from Guinea), L. angolensis (8 from Cameroon, 4 from Guinea), M. pusillus (2 from the DRC, 1 from Guinea), R. aegyptiacus (45 from Cameroon, 40 from Guinea), E. gambianus (25 from Guinea), and Mops sp. (20 from Cameroon), all were negative for Zaire Ebola virus RNA.
To clarify the role of bats in Ebola virus ecology and identify where the virus circulates between outbreaks, we tested >4,000 bats, almost doubling the total number of samples tested in all previous studies in Africa (5–7,42). We provided data on bats from Cameroon, added to the existing data on bats from Guinea and the DRC, and substantially increased the data available on insectivorous bats. We tested samples with the same assay, enabling comparison across species and countries. We used different statistical methods to determine positive sample numbers and expressed the proportion of reactive samples as a range on the basis of the different cutoff values proposed by those methods. As has been done in studies of human Zaire Ebola virus survivors (28,43), we defined Zaire and Sudan Ebola virus positivity as the presence of antibodies to both nucleoprotein and glycoprotein. As such, we estimated that 2–37 (0.05%–0.92%) bats were seropositive for Zaire Ebola virus and 0–30 (0%–0.75%) bats were seropositive for Sudan Ebola virus (Table 3). Among insectivorous bats, we observed Zaire and Sudan Ebola virus antibodies only in Mops sp. bats, an observation that has previously been observed (13). We provided information on insectivorous Miniopterus and Rhinolophus bats and extended knowledge on Mops and Hipposideros bats; all 1,200 Hipposideros samples were seronegative. We confirmed the presence of Zaire Ebola virus antibodies in only 1 of 3 frugivorous species in which Zaire Ebola virus RNA has been reported, that is, in H. monstrosus but not E. franqueti or M. torquata bats (6). However, this result might have been influenced by sample size, test used, and interpretation criteria. We confirmed antibodies in E. helvum bats and showed that Zaire Ebola virus antibodies are widespread among this species across Africa: Ghana and Zambia, and with our data, also Cameroon, Guinea, and the DRC (13,14,16). We confirmed antibodies in R. aegyptiacus bats from Cameroon and Guinea, in agreement with previous findings in these bats from the Congo and Gabon (13). For E. gambianus bats from Ghana, we also observed Zaire Ebola virus reactivity of samples from this species in Guinea (15). In contrast with a previous study, we observed Sudan Ebola virus antibodies (not Zaire Ebola virus antibodies) in M. pusillus bats (13). We also identified Zaire Ebola virus antibodies in L. angolensis bats from Cameroon, although only when using less stringent cutoff calculations.
When combining data from previous Zaire Ebola virus seroprevalence studies in bats with data from our study, only 1 insectivorous bat species (Mops sp.) and 8 frugivorous bat species (E. helvum, E. gambianus, E. franqueti, H. monstrosus, L. angolensis, M. pusillus, M. torquata, R. aegyptiacus) exhibited Zaire Ebola virus antibodies (13–16). As seen in bat samples from Zambia, we observed in this study Sudan Ebola virus antibodies in E. helvum bats from Guinea, Cameroon, and the DRC, suggesting that Zaire and Sudan Ebola viruses co-circulate and could be widespread among this species. However, only 1 other study has tested for Ebola viruses other than Zaire Ebola virus in E. helvum bats (16). In our study, we also observed Sudan Ebola virus antibodies in Mops sp., H. monstrosus, and R. aegyptiacus bats in Cameroon. Almost all samples were positive for either Zaire or Sudan Ebola virus but not for both.
Despite the presence of Ebola virus antibodies, the role of bats as reservoir species remains unclear because viral RNA detection is rare. In only 1 study Zaire Ebola virus RNA was amplified in a few bats (6). Thus, antibodies might reflect previous acute infection with viral clearance. Unlike inoculations with Marburg virus (44–46), experimental inoculation of R. aegyptiacus bats with Zaire Ebola virus leads to antibody development but infrequent or absent detection of viral RNA or shedding (44,47). R. aegyptiacus bats are therefore able to clear Zaire Ebola virus after a short infectious period without viral shedding and with little or no transmission. No antibodies or viral RNA were detected in noninoculated bats housed with experimentally Zaire Ebola virus–infected bats (44). Whether this low level of infectiousness also occurs for other bat species that carry Ebola virus antibodies remains to be determined. Zaire Ebola virus was experimentally inoculated in other bat species (M. condylurus, Chaerephon pumilus, and Epomophorus wahlbergi) in only 1 study; virus replication was seen in all species, and fecal shedding was seen in E. wahlbergi bats (48). R. aegyptiacus bats experimentally infected with Marburg virus were shown to develop antibodies that protect against reinfection (49). Long-term survival with Zaire Ebola virus antibodies has been reported with E. helvum bats from Ghana but without information on protection (14). Among insectivorous bats, the presence of Ebola virus antibodies in only Mops sp. is striking, suggesting higher exposure or susceptibility compared with other insectivorous bats.
In conclusion, we demonstrated higher rates of Ebola virus antibodies in frugivorous than in insectivorous bats. The total number of frugivorous species shown to be Zaire Ebola virus seropositive has increased to 8, and 1 insectivorous bat species (Mops sp.) was confirmed to be seropositive. Zaire and Sudan Ebola viruses circulate in different species across Africa, with potential co-circulation of both viruses in some species. Although we have data on >8,000 bats from >40 species, this sample size is small, given the high numbers of bats that constitute colonies. This study illustrates the complexity of tracking the animal reservoir of Ebola viruses, not only because sampling of wild bats without performing euthanasia is difficult and time-consuming but also because of the absence of a reference standard for serologic tests. To clarify the significance of Ebola virus antibodies, documenting the extent to which viral RNA and shedding can be detected in species with antibodies is crucial for predicting and controlling the risk for new outbreaks. Efforts must continue not only to sample bats but also other animals to elucidate where the virus circulates in wildlife.
Drs. Mbala Kingebeni and Keita are researchers from ASTRE of Centre de coopération internationale en recherche agronomique pour le développement, Institut national de la recherche agronomique and Univerisity of Montpellier, Montpellier, France. Their research interests include characterization of hosts that harbor zoonotic pathogens.
Acknowledgments
We thank the staffs from the Ministry of Health and Ministry of Environment and the national ethics committees from the DRC, Cameroon, and Guinea for permission to perform this study. We thank all the field staff from the DRC (Guy Midingi and Servet Kimbonza); Guinea (Souana Goumou, Mamadou Kalif Diallo, Pépé Justin Beavogui, Philippe Kolié, Michel Guilavogui); and Cameroon (Innocent Ndong Bass, Aime Mebenga, Joseph Moudindo, Thomas Atemkem) for the collection of bat samples. We thank the staffs of the National Institute of Biomedical Research (Kinshasa, the DRC), the Kongo Central Provincial Government (Matadi, the DRC), and Projet PRESICA; Donald Mbohli from Project Grand Singes; and the staff of the Institut National de Santé Publique (Conakry, Guinea) for logistical support in the field. We thank Seny Mane for his involvement and support in the implementation of this project and Daouda Bangoura for his constant support for this project and the facilitation of field missions. We thank the veterinary staff from the Parc Zoologique de Montpellier and Wilhelma Zoo and Botanical Garden for providing control samples.
This work was supported in part by grants from Institut National de la Santé et de la Recherche Médicale, the Ebola Task Force, REACTing, EBO-SURSY project funded by the European Union, Institut de Recherche pour le Développement (IRD), and Christophe Mérieux Prize 2015 (to J.-J.M.T.). A.K.K. was supported by a fellowship from the IRD and the University of Montpellier (MUSE, ANR-16-IDEX-0006). C.-J.V.-A. was supported by a fellowship from IRD, Labex EpiGenMed via the National Research Agency, Programme for Future Investment (ANR-10-LABX-12-01), and the University of Montpellier.
References
- Mylne A, Brady OJ, Huang Z, Pigott DM, Golding N, Kraemer MU, et al. A comprehensive database of the geographic spread of past human Ebola outbreaks. Sci Data. 2014;1:140042. DOIPubMedGoogle Scholar
- Baize S, Pannetier D, Oestereich L, Rieger T, Koivogui L, Magassouba N, et al. Emergence of Zaire Ebola virus disease in Guinea. N Engl J Med. 2014;371:1418–25. DOIPubMedGoogle Scholar
- Maganga GD, Kapetshi J, Berthet N, Kebela Ilunga B, Kabange F, Mbala Kingebeni P, et al. Ebola virus disease in the Democratic Republic of Congo. N Engl J Med. 2014;371:2083–91. DOIPubMedGoogle Scholar
- World Health Organization. Ebola outbreak Democratic Republic of the Congo 2017. 2017 Jul 2 [cited 2018 May 3]. http://www.who.int/emergencies/ebola-DRC-2017/en/
- Pigott DM, Millear AI, Earl L, Morozoff C, Han BA, Shearer FM, et al. Updates to the zoonotic niche map of Ebola virus disease in Africa. eLife. 2016;5:e16412. DOIPubMedGoogle Scholar
- Leroy EM, Kumulungui B, Pourrut X, Rouquet P, Hassanin A, Yaba P, et al. Fruit bats as reservoirs of Ebola virus. Nature. 2005;438:575–6. DOIPubMedGoogle Scholar
- Leendertz SA, Gogarten JF, Düx A, Calvignac-Spencer S, Leendertz FH. Assessing the evidence supporting fruit bats as the primary reservoirs for Ebola viruses. EcoHealth. 2016;13:18–25. DOIPubMedGoogle Scholar
- Kamins AO, Rowcliffe JM, Ntiamoa-Baidu Y, Cunningham AA, Wood JL, Restif O. Characteristics and risk perceptions of Ghanaians potentially exposed to bat-borne zoonoses through bushmeat. EcoHealth. 2015;12:104–20. DOIPubMedGoogle Scholar
- Gonzalez JP, Pourrut X, Leroy E. Ebolavirus and other filoviruses. Curr Top Microbiol Immunol. 2007;315:363–87. DOIPubMedGoogle Scholar
- Leroy EM, Epelboin A, Mondonge V, Pourrut X, Gonzalez JP, Muyembe-Tamfum JJ, et al. Human Ebola outbreak resulting from direct exposure to fruit bats in Luebo, Democratic Republic of Congo, 2007. Vector Borne Zoonotic Dis. 2009;9:723–8. DOIPubMedGoogle Scholar
- Marí Saéz A, Weiss S, Nowak K, Lapeyre V, Zimmermann F, Düx A, et al. Investigating the zoonotic origin of the West African Ebola epidemic. EMBO Mol Med. 2015;7:17–23. DOIPubMedGoogle Scholar
- Pourrut X, Délicat A, Rollin PE, Ksiazek TG, Gonzalez JP, Leroy EM. Spatial and temporal patterns of Zaire ebolavirus antibody prevalence in the possible reservoir bat species. J Infect Dis. 2007;196(Suppl 2):S176–83. DOIPubMedGoogle Scholar
- Pourrut X, Souris M, Towner JS, Rollin PE, Nichol ST, Gonzalez JP, et al. Large serological survey showing cocirculation of Ebola and Marburg viruses in Gabonese bat populations, and a high seroprevalence of both viruses in Rousettus aegyptiacus. BMC Infect Dis. 2009;9:159. DOIPubMedGoogle Scholar
- Hayman DT, Emmerich P, Yu M, Wang LF, Suu-Ire R, Fooks AR, et al. Long-term survival of an urban fruit bat seropositive for Ebola and Lagos bat viruses. PLoS One. 2010;5:e11978. DOIPubMedGoogle Scholar
- Hayman DT, Yu M, Crameri G, Wang LF, Suu-Ire R, Wood JL, et al. Ebola virus antibodies in fruit bats, Ghana, West Africa. Emerg Infect Dis. 2012;18:1207–9. DOIPubMedGoogle Scholar
- Ogawa H, Miyamoto H, Nakayama E, Yoshida R, Nakamura I, Sawa H, et al. Seroepidemiological prevalence of multiple species of filoviruses in fruit bats (Eidolon helvum) migrating in Africa. J Infect Dis. 2015;212(Suppl 2):S101–8. DOIPubMedGoogle Scholar
- Towner JS, Pourrut X, Albariño CG, Nkogue CN, Bird BH, Grard G, et al. Marburg virus infection detected in a common African bat. PLoS One. 2007;2:e764. DOIPubMedGoogle Scholar
- Towner JS, Amman BR, Sealy TK, Carroll SA, Comer JA, Kemp A, et al. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Pathog. 2009;5:e1000536. DOIPubMedGoogle Scholar
- Kuzmin IV, Niezgoda M, Franka R, Agwanda B, Markotter W, Breiman RF, et al. Marburg virus in fruit bat, Kenya. Emerg Infect Dis. 2010;16:352–4. DOIPubMedGoogle Scholar
- Swanepoel R, Smit SB, Rollin PE, Formenty P, Leman PA, Kemp A, et al.; International Scientific and Technical Committee for Marburg Hemorrhagic Fever Control in the Democratic Republic of Congo. Studies of reservoir hosts for Marburg virus. Emerg Infect Dis. 2007;13:1847–51. DOIPubMedGoogle Scholar
- Negredo A, Palacios G, Vázquez-Morón S, González F, Dopazo H, Molero F, et al. Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog. 2011;7:e1002304. DOIPubMedGoogle Scholar
- Yang XL, Zhang YZ, Jiang RD, Guo H, Zhang W, Li B, et al. Genetically diverse filoviruses in Rousettus and Eonycteris spp. bats, China, 2009 and 2015. Emerg Infect Dis. 2017;23:482–6. DOIPubMedGoogle Scholar
- World Health Organization. Ebola situation report - 30 March 2016. 2016 [cited 2018 May 3].http://apps.who.int./ebola/current-situation/ebola-situation-report-30-march-2016
- Dudas G, Carvalho LM, Bedford T, Tatem AJ, Baele G, Faria NR, et al. Virus genomes reveal factors that spread and sustained the Ebola epidemic. Nature. 2017;544:309–15. DOIPubMedGoogle Scholar
- Pigott DM, Deshpande A, Letourneau I, Morozoff C, Reiner RC Jr, Kraemer MUG, et al. Local, national, and regional viral haemorrhagic fever pandemic potential in Africa: a multistage analysis. Lancet. 2017;390:2662–72. DOIPubMedGoogle Scholar
- Breman JG, Johnson KM, van der Groen G, Robbins CB, Szczeniowski MV, Ruti K, et al.; Ebola Virus Study Teams. A search for Ebola virus in animals in the Democratic Republic of the Congo and Cameroon: ecologic, virologic, and serologic surveys, 1979-1980. J Infect Dis. 1999;179(Suppl 1):S139–47. DOIPubMedGoogle Scholar
- Leirs H, Mills JN, Krebs JW, Childs JE, Akaibe D, Woollen N, et al. Search for the Ebola virus reservoir in Kikwit, Democratic Republic of the Congo: reflections on a vertebrate collection. J Infect Dis. 1999;179(Suppl 1):S155–63. DOIPubMedGoogle Scholar
- Ayouba A, Touré A, Butel C, Keita AK, Binetruy F, Sow MS, et al. Development of a sensitive and specific serological assay based on luminex technology for detection of antibodies to Zaire Ebola virus. J Clin Microbiol. 2016;55:165–76. DOIPubMedGoogle Scholar
- Peel AJ, McKinley TJ, Baker KS, Barr JA, Crameri G, Hayman DT, et al. Use of cross-reactive serological assays for detecting novel pathogens in wildlife: assessing an appropriate cutoff for henipavirus assays in African bats. J Virol Methods. 2013;193:295–303. DOIPubMedGoogle Scholar
- Gilbert AT, Fooks AR, Hayman DT, Horton DL, Müller T, Plowright R, et al. Deciphering serology to understand the ecology of infectious diseases in wildlife. EcoHealth. 2013;10:298–313. DOIPubMedGoogle Scholar
- Lardeux F, Torrico G, Aliaga C. Calculation of the ELISA’s cut-off based on the change-point analysis method for detection of Trypanosoma cruzi infection in Bolivian dogs in the absence of controls. Mem Inst Oswaldo Cruz. 2016;111:501–4. DOIPubMedGoogle Scholar
- Killick R, Eckley IA. changepoint: an R package for changepoint analysis. J Stat Softw. 2014;58:1–19. DOIGoogle Scholar
- Hinkley DV. Inference about the change-point in a sequence of random variables. Biometrika. 1970;57:1–17. DOIGoogle Scholar
- Laing ED, Mendenhall IH, Linster M, Low DHW, Chen Y, Yan L, et al. Serologic evidence of fruit bat exposure to filoviruses, Singapore, 2011–2016. Emerg Infect Dis. 2018;24:114–7. DOIPubMedGoogle Scholar
- Cullen AC, Frey HC. Probabilistic techniques in exposure assessment. New York: Plenum Press; 1999. p. 81–159.
- Delignette-Muller ML, Dutang C. fitdistrplus: an R package for fitting distributions. J Stat Softw. 2015;64:1–34. DOIGoogle Scholar
- Monleau M, Montavon C, Laurent C, Segondy M, Montes B, Delaporte E, et al. Evaluation of different RNA extraction methods and storage conditions of dried plasma or blood spots for human immunodeficiency virus type 1 RNA quantification and PCR amplification for drug resistance testing. J Clin Microbiol. 2009;47:1107–18. DOIPubMedGoogle Scholar
- Guichet E, Serrano L, Laurent C, Eymard-Duvernay S, Kuaban C, Vidal L, et al. Comparison of different nucleic acid preparation methods to improve specific HIV-1 RNA isolation for viral load testing on dried blood spots. J Virol Methods. 2018;251:75–9. DOIPubMedGoogle Scholar
- Irwin DM, Kocher TD, Wilson AC. Evolution of the cytochrome b gene of mammals. J Mol Evol. 1991;32:128–44. DOIPubMedGoogle Scholar
- Kocher TD, Thomas WK, Meyer A, Edwards SV, Pääbo S, Villablanca FX, et al. Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci U S A. 1989;86:6196–200. DOIPubMedGoogle Scholar
- He B, Feng Y, Zhang H, Xu L, Yang W, Zhang Y, et al. Filovirus RNA in fruit bats, China. Emerg Infect Dis. 2015;21:1675–7. DOIPubMedGoogle Scholar
- Han BA, Schmidt JP, Alexander LW, Bowden SE, Hayman DT, Drake JM. Undiscovered bat hosts of filoviruses. PLoS Negl Trop Dis. 2016;10:e0004815. DOIPubMedGoogle Scholar
- Rimoin AW, Lu K, Bramble MS, Steffen I, Doshi RH, Hoff NA, et al. Ebola virus neutralizing antibodies detectable in survivors of theYambuku, Zaire outbreak 40 years after infection. J Infect Dis. 2018;217:223–31. DOIPubMedGoogle Scholar
- Paweska JT, Storm N, Grobbelaar AA, Markotter W, Kemp A, Jansen van Vuren P. Experimental inoculation of Egyptian fruit bats (Rousettus aegyptiacus) with Ebola virus. Viruses. 2016;8:29. DOIPubMedGoogle Scholar
- Amman BR, Jones ME, Sealy TK, Uebelhoer LS, Schuh AJ, Bird BH, et al. Oral shedding of Marburg virus in experimentally infected Egyptian fruit bats (Rousettus aegyptiacus). J Wildl Dis. 2015;51:113–24. DOIPubMedGoogle Scholar
- Schuh AJ, Amman BR, Jones ME, Sealy TK, Uebelhoer LS, Spengler JR, et al. Modelling filovirus maintenance in nature by experimental transmission of Marburg virus between Egyptian rousette bats. Nat Commun. 2017;8:14446. DOIPubMedGoogle Scholar
- Jones ME, Schuh AJ, Amman BR, Sealy TK, Zaki SR, Nichol ST, et al. Experimental inoculation of Egyptian rousette bats (Rousettus aegyptiacus) with viruses of the Ebolavirus and Marburgvirus genera. Viruses. 2015;7:3420–42. DOIPubMedGoogle Scholar
- Swanepoel R, Leman PA, Burt FJ, Zachariades NA, Braack LE, Ksiazek TG, et al. Experimental inoculation of plants and animals with Ebola virus. Emerg Infect Dis. 1996;2:321–5. DOIPubMedGoogle Scholar
- Storm N, Jansen Van Vuren P, Markotter W, Paweska JT. Antibody responses to Marburg virus in Egyptian rousette bats and their role in protection against infection. Viruses. 2018;10:73. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: November 20, 2018
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
Table of Contents – Volume 24, Number 12—December 2018
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Adress for correspondence: Martine Peeters, TransVIHMI, Institut de Recherche pour le Développement, 911 Ave Agropolis, BP34394 Montpellier CEDEX 1, France
Top