Volume 28, Number 8—August 2022
Dispatch
Association of Phylogenomic Relatedness among Neisseria gonorrhoeae Strains with Antimicrobial Resistance, Austria, 2016–2020
Cite This Article
Citation for Media
Abstract
We investigated genomic determinants of antimicrobial resistance in 1,318 Neisseria gonorrhoeae strains isolated in Austria during 2016–2020. Sequence type (ST) 9363 and ST11422 isolates had high rates of azithromycin resistance, and ST7363 isolates correlated with cephalosporin resistance. These results underline the benefit of genomic surveillance for antimicrobial resistance monitoring.
Gonorrhea, a sexually transmissible infection (STI) caused by Neisseria gonorrhoeae, is the second most common bacterial STI (1). Most gonorrhea cases are mild, but serious complications can occur. Gonorrhea is treated with antibiotics, and the recommended treatment is dual extended-spectrum cephalosporin (ESC)/azithromycin therapy or ceftriaxone monotherapy (2).
One of the main characteristics of N. gonorrhoeae is the plasticity of its genome, favoring the acquisition and dispersion of antimicrobial resistance (AMR). AMR is an increasing issue for gonorrhea treatment, and untreatable gonorrhea represents an imminent global health threat (3).
Whole-genome sequencing (WGS) provides high-resolution data that can support AMR surveillance. We combined phenotypic AMR testing with WGS to investigate 1,318 N. gonorrhoeae strains isolated in Austria during 2016–2020 and identify genetic risk factors associated with AMR.
This study encompassed 1,318 N. gonorrhoeae isolates collected in Austria during 2016–2020; isolates were available at the National Reference Centre for Gonococci. We tested all isolates for phenotypic resistance to azithromycin, cefixime, ceftriaxone, ciprofloxacin, tetracycline, and benzylpenicillin, as well as production of β-lactamase (i.e., cefinase positive) (Appendix). We followed European Committee on Antimicrobial Susceptibility Testing guidelines (4) to determine MIC thresholds used in this study.
We performed genomic DNA isolation, WGS, assembly, and contig filtering as described previously (5) (Appendix). We deposited raw reads in the National Center for Biotechnology Information Sequence Read Archive (project no. PRJNA771206). We obtained sequences types (STs) from WGS data by using the PubMLST schemes (6,7). We generated a local N. gonorrhoeae core-genome multilocus sequence typing (cgMLST) scheme with SeqSphere+ target definer tool version 6.0.0 (Ridom, ttps://www.ridom.de) (5) (Appendix). We investigated AMR genes by using allele libraries based on PathogenWatch in TOML format version 0.0.14 (8).
We performed time series analysis, linear regression, univariate analysis, multivariate analysis (logistic regression), and data visualization by using R version 4.0.4 (Appendix). We defined statistical significance as p<0.05. We computed neighbor-joining trees in SeqSphere+ by using the number of cgMLST allelic differences and exported the trees into R.
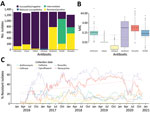
We classified isolates according to AMR (Figure 1, panel A; Table) and determined MIC distributions (Figure 1, panel B). We observed high levels of resistance to ciprofloxacin (60%) and tetracycline (46%) (Figure 1, panel A), which increased 5% per year for ciprofloxacin (p<0.0001) and 6% per year for tetracycline (p<0.0001). The percentage of penicillin-resistant isolates was 16% and decreased over the study period (2% per year; p<0.0001) (Figure 1, panel C); 14% of isolates were cefinase-positive, which increased by 2.7% per year (p<0.0001).
We detected azithromycin resistance in 9% of the isolates, which increased by 5% per year (p<0.0001) (Figure 1). Two isolates from 2020 exhibited high levels of azithromycin resistance (MIC >256 µg/mL) but no other AMR. Resistance to ESC was rare; only 3% of isolates were resistant to cefixime, none were resistant to ceftriaxone, and 2.5% had reduced susceptibility to ceftriaxone (MIC >0.032 µg/mL). Cefixime resistance decreased by 0.9% per year (p<0.0001). Among cefixime-resistant isolates, 23/35 were resistant to ciprofloxacin and penicillin, qualifying as multidrug resistant.
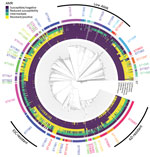
The isolates belonged to 119 different STs in mutlilocus sequence typing, including 23 newly defined (STs 15803–15825). The most prevalent STs were ST7363 (170 isolates), ST9363 (151 isolates), and ST8156 (113 isolates), which comprised 33% of the isolates. We identified 215 NG-MAST types for 873/1,318 isolates; the most prevalent STs were 12302 (73 isolates), 5441 (59 isolates), and 387 (50 isolates). cgMLST showed a branch including isolates with no or little AMR (Figure 2). We found no clear correlation with the cgMLST classification for penicillin, cefinase, tetracycline, and ciprofloxacin resistance. All cefixime-resistant isolates belonged to a single branch of ST7363 isolates, which also contained 24/32 isolates with reduced susceptibility to ceftriaxone. This branch had above average rates of ciprofloxacin, tetracycline, and penicillin resistance. A branch containing ST9363 and ST11422 isolates had a high rate of azithromycin resistance.
We searched isolate sequences for genes and point mutations associated with AMR (Appendix Table 3). For ciprofloxacin resistance, gyrA D95 substitutions were the main risk factor (adjusted odds ratio [aOR] 7.56 [95% CI 2.33–33.1]) and explained >99% of ciprofloxacin resistance. Tetracycline resistance was strongly associated with tetM carriage (aOR 157 [95% CI 48–965]), which we found in 33% of tetracycline-resistant isolates. For β-lactams, the main risk factor was blaTEM carriage (aOR 67.9 [95% CI 35.2–139] for penicillin and aOR 234 [95% CI 93.3–683] for cefinase). Mutations in penA were also associated with cefinase positivity (aOR 35.6 [95% CI 14–97.4]).
We found mutations in the macAB promoter or mosaic mtr genes in 138/149 azithromycin-resistant isolates (93%). All cefixime-resistant isolates carried penA G545S substitution. The major risk factor for reduced susceptibility to ceftriaxone was penA A501T/V (aOR 73.9 [95% CI 6.9–3,170]).
This study combined phenotypic AMR and genomic data to analyze N. gonorrhoeae strains isolated in Austria during 2016–2020. We used a convenience sample (National Reference Centre for Gonococci collection) and results should be interpreted in light of this limitation. The percentage of N. gonorrhoeae strains resistant to azithromycin, ciprofloxacin, and tetracycline, or producing β-lactamase was increasing during the study period. The rate of azithromycin resistance rate was >13% during 2019–2020, which was high considering that an azithromycin/cefixime combination is a standard treatment for gonorrhea (2). We found no ceftriaxone-resistant isolates, and cefixime resistance rate was low.
We performed isolate typing by using multilocus sequence typing, N. gonorrhoeae multiantigen sequence typing (NG-MAST), and cgMLST. Only 37 isolates belonged to ST1901, which was predominant in isolates from Austria in a European study in 2013, highlighting the fast diversification of N. gonorrhoeae (9). The most common NG-MAST type was 12302; all isolates belonged to ST9363 and 71% were resistant to azithromycin. NG-MAST type 12302 and ST9363 have been associated with azithromycin resistance in other studies (10,11). cgMLST classification highlighted 3 branches with specific AMR patterns: 1 with low rates of AMR, 1 including azithromycin-resistant isolates, and 1 including ESC-resistant isolates. Previous studies comparing AMR and phylogenomic distributions in different countries showed either that azithromycin/ESC resistance emerged repeatedly in different networks or that their spread was largely clonal (12,13). In Austria, azithromycin and ESC resistance clustering was in favor of single introductions. The use of cgMLST among available classification methods has limitations (i.e., no counting of mutations within 1 gene, exclusion of intergenic regions, and resolution) but also advantages (i.e., no correction of recombination events necessary and one scheme fitting all isolates). This tool corresponds to the need for surveillance, where its lower resolution does not have a major effect.
We used our WGS data to search for genetic determinants of AMR (8,14). Ciprofloxacin resistance matched well with gyrA mutations (9,12). Tetracycline resistance correlated with tetM, and penicillin resistance correlated blaTEM. Mutations in penA and mtrR were associated with ESC resistance. Neither substitution C1192U in 16S rDNA nor rpsE V25 mutations, associated with spectinomycin resistance, were found, suggesting a low prevalence of spectinomycin resistance.
Our study provides an overview of the N. gonorrhoeae strains circulating in Austria and their evolution over the past 5 years, both at the phenotypic and genomic level. It also underlines the benefits of genomic surveillance of N. gonorrhoeae, which can support epidemiologic investigations and provide information on specific genes and alleles thought to confer AMR (14).
Dr. Schaeffer is a public health microbiologist at the Austrian Agency for Health and Food Safety, Vienna, Austria. Her research interests include emerging pathogens, severe human diseases, and genomic analysis.
Acknowledgments
We thank the staff of the Gonococci National Reference Centre and other partner institutions for the collection, isolation, and characterization of the study isolates. We thank Loredana Ingrosso for her feedback throughout the project and on the manuscript.
All co-authors in this manuscript declare that no funding was received from any funding agency in the public, commercial, or not-for-profit sectors. J.S. was supported by a grant from the European Public Health Microbiology Training Programme, European Centre for Disease Prevention and Control (grant agreement no. 1 ECD.7550, implementing ECDC/GRANT/2017/003).
References
- Rowley J, Vander Hoorn S, Korenromp E, Low N, Unemo M, Abu-Raddad LJ, et al. Chlamydia, gonorrhoea, trichomoniasis and syphilis: global prevalence and incidence estimates, 2016. Bull World Health Organ. 2019;97:548–562P. DOIPubMedGoogle Scholar
- World Health Organization. WHO guidelines for the treatment of Neisseria gonorrhoeae. Report no. 978–92–4-154969–1. 2016 [cited 2022 Jan 8]. https://apps.who.int/iris/bitstream/handle/10665/246114/9789241549691-eng.pdf
- World Health Organization. Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics. 2017 [cited 2022 Jan 8]. https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters, version 11.0. 2021 [cited 2022 Jan 8]. https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_10.0_Breakpoint_Tables.pdf
- Hirk S, Lepuschitz S, Cabal Rosel A, Huhulescu S, Blaschitz M, Stöger A, et al. Draft genome sequences of interpatient and intrapatient epidemiologically linked Neisseria gonorrhoeae isolates. Genome Announc. 2018;6:e00319–18. DOIPubMedGoogle Scholar
- Maiden MC, Bygraves JA, Feil E, Morelli G, Russell JE, Urwin R, et al. Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci U S A. 1998;95:3140–5. DOIPubMedGoogle Scholar
- Martin IM, Ison CA, Aanensen DM, Fenton KA, Spratt BG. Rapid sequence-based identification of gonococcal transmission clusters in a large metropolitan area. J Infect Dis. 2004;189:1497–505. DOIPubMedGoogle Scholar
- Sánchez-Busó L, Yeats CA, Taylor B, Goater RJ, Underwood A, Abudahab K, et al. A community-driven resource for genomic epidemiology and antimicrobial resistance prediction of Neisseria gonorrhoeae at Pathogenwatch. Genome Med. 2021;13:61. DOIPubMedGoogle Scholar
- Harris SR, Cole MJ, Spiteri G, Sánchez-Busó L, Golparian D, Jacobsson S, et al.; Euro-GASP study group. Public health surveillance of multidrug-resistant clones of Neisseria gonorrhoeae in Europe: a genomic survey. Lancet Infect Dis. 2018;18:758–68. DOIPubMedGoogle Scholar
- Sawatzky P, Demczuk W, Lefebvre B, Allen V, Diggle M, Hoang L, et al. Increasing azithromycin resistance in Neisseria gonorrhoeae due to NG-MAST 12302 clonal spread in Canada, 2015 to 2018. Antimicrob Agents Chemother. 2022;66:
e0168821 . DOIPubMedGoogle Scholar - Williamson DA, Chow EPF, Gorrie CL, Seemann T, Ingle DJ, Higgins N, et al. Bridging of Neisseria gonorrhoeae lineages across sexual networks in the HIV pre-exposure prophylaxis era. Nat Commun. 2019;10:3988. DOIPubMedGoogle Scholar
- Lee RS, Seemann T, Heffernan H, Kwong JC, Gonçalves da Silva A, Carter GP, et al. Genomic epidemiology and antimicrobial resistance of Neisseria gonorrhoeae in New Zealand. J Antimicrob Chemother. 2018;73:353–64. DOIPubMedGoogle Scholar
- Harrison OB, Cehovin A, Skett J, Jolley KA, Massari P, Genco CA, et al. Neisseria gonorrhoeae population genomics: use of the gonococcal core genome to improve surveillance of antimicrobial resistance. J Infect Dis. 2020;222:1816–25. DOIPubMedGoogle Scholar
- Demczuk W, Martin I, Sawatzky P, Allen V, Lefebvre B, Hoang L, et al. Equations to predict antimicrobial MICs in Neisseria gonorrhoeae using molecular antimicrobial resistance determinants. Antimicrob Agents Chemother. 2020;64:e02005–19. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: July 11, 2022
Table of Contents – Volume 28, Number 8—August 2022
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Justine Schaeffer, Österreichische Agentur für Gesundheit und Ernährungssicherheit, Währinger Straße 25A, 1096 Vienna, Austria
Top