Metagenomic Identification of Fusarium solani Strain as Cause of US Fungal Meningitis Outbreak Associated with Surgical Procedures in Mexico, 2023
Charles Y. Chiu

, Venice Servellita, Mikael de Lorenzi-Tognon, Patrick Benoit, Nanami Sumimoto, Abiodun Foresythe, Filipe M. Cerqueira, Natalie Williams-Bouyer, Ping Ren, Lauren Nicholas S. Herrera, David C. Gaston, Leanna Sayyad, Shannon L. Whitmer, John Klena, Holenarasipur R. Vikram, Jeremy A.W. Gold, Lalitha Gade, Lindsay Parnell, Elizabeth Misas, Tom M. Chiller, Isabel S. Griffin, Sridhar V. Basavaraju, Dallas J. Smith, Anastasia P. Litvintseva, and Nancy A. Chow
Author affiliation: Chan-Zuckerberg Biohub, San Francisco, California, USA (C.Y. Chiu); University of California San Francisco, San Francisco (C.Y. Chiu, V. Servellita, M. de Lorenzi-Tognon, P. Benoit, N. Sumimoto, A. Foresythe); Abbott Pandemic Defense Coalition, Abbott Park, Illinois, USA (C.Y. Chiu, V. Servellita, M. de Lorenzi-Tognon, P. Benoit, N. Sumimoto, A. Foresythe); The University of Texas Medical Branch at Galveston, Galveston, Texas, USA (F.M. Cerqueira, N. Williams-Bouyer, P. Ren); Vanderbilt University Medical Center, Nashville, Tennessee, USA (L.N.S. Herrera, D.C. Gaston); Centers for Disease Control and Prevention, Atlanta, Georgia, USA (L. Sayyad, S.L. Whitmer, J. Klena, J.A.W. Gold, L. Gade, L. Parnell, E. Misas, T.M. Chiller, I.S. Griffin, S.V. Basavaraju, D.J. Smith, A.P. Litvintseva, N.A. Chow); Mayo Clinic, Phoenix, Arizona, USA (H.R. Vikram)
Main Article
Figure 2
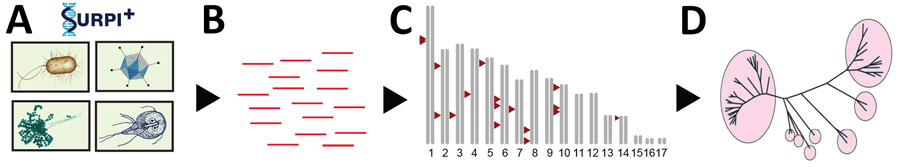
Figure 2. Flowchart showing the metaMELT analysis workflow used for identification of Fusarium solani strain as cause of fungal meningitis US outbreak associated with surgical procedures in Mexico, 2023. metaMELT (metagenomic multiple extended locus typing), is a novel analytic technique for simultaneously diagnosing the infection and characterizing the interrelatedness of F. solani strains. metaMELT uses the following steps: A) perform mNGS analysis of patient samples (i.e., cerebrospinal fluid, plasma, or brain tissue), using the SURPI+ computational pipeline (https://github.com/chiulab/SURPI-plus-dist) to identify pathogens; B) identify F. solani reads; C) map reads to the F. solani reference genome and then extract and concatenate; D) perform phylogenetic analysis on concatenated sequences. SURPI, sequence-based ultra-rapid pathogen identification.
Main Article
Page created: March 21, 2025
Page updated: April 25, 2025
Page reviewed: April 25, 2025
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
