Volume 31, Number 5—May 2025
Research
Metagenomic Identification of Fusarium solani Strain as Cause of US Fungal Meningitis Outbreak Associated with Surgical Procedures in Mexico, 2023
Figure 3
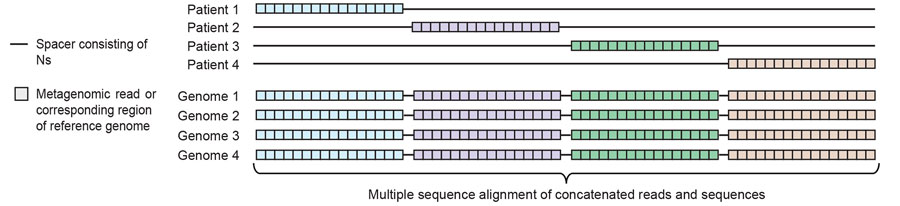
Figure 3. Diagram of concatenation and alignment step of the metaMELT procedure (metagenomic multiple extended locus typing, a novel analytic technique for simultaneously diagnosing the infection and characterizing the interrelatedness of Fusarium solani strains) used for identification of F. solani strain as cause of fungal meningitis US outbreak associated with surgical procedures in Mexico, 2023. The diagram shows concatenated metagenomic next-generation sequencing reads from 4 patients and the corresponding regions extracted from reference genomes, which are aligned to by using MAFFT version 7.388 (22). This diagram demonstrates the steps shown in Figure 2, panels C–D.
References
- Popovich KJ, Snitkin ES. Whole genome sequencing—implications for infection prevention and outbreak investigations. Curr Infect Dis Rep. 2017;19:15.PubMedGoogle Scholar
- Gilchrist CA, Turner SD, Riley MF, Petri WA Jr, Hewlett EL. Whole-genome sequencing in outbreak analysis. Clin Microbiol Rev. 2015;28:541–63. DOIPubMedGoogle Scholar
- Cuomo CA. Harnessing whole genome sequencing in medical mycology. Curr Fungal Infect Rep. 2017;11:52–9.PubMedGoogle Scholar
- Maiden MC, Jansen van Rensburg MJ, Bray JE, Earle SG, Ford SA, Jolley KA, et al. MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol. 2013;11:728–36. DOIPubMedGoogle Scholar
- Urwin R, Maiden MC. Multi-locus sequence typing: a tool for global epidemiology. Trends Microbiol. 2003;11:479–87.PubMedGoogle Scholar
- Benoit P, Brazer N, de Lorenzi-Tognon M, Kelly E, Servellita V, Oseguera M, et al. Seven-year performance of a clinical metagenomic next-generation sequencing test for diagnosis of central nervous system infections. Nat Med. 2024;30:3522–33.PubMedGoogle Scholar
- Bergin SP, Chemaly RF, Dadwal SS, Hill JA, Lee YJ, Haidar G, et al. Plasma microbial cell-free DNA sequencing in immunocompromised patients with pneumonia: a prospective observational study. Clin Infect Dis. 2024;78:775–84.PubMedGoogle Scholar
- Blauwkamp TA, Thair S, Rosen MJ, Blair L, Lindner MS, Vilfan ID, et al. Analytical and clinical validation of a microbial cell-free DNA sequencing test for infectious disease. Nat Microbiol. 2019;4:663–74.PubMedGoogle Scholar
- Fourgeaud J, Regnault B, Ok V, Da Rocha N, Sitterlé É, Mekouar M, et al. Performance of clinical metagenomics in France: a prospective observational study. Lancet Microbe. 2024;5:e52–61.PubMedGoogle Scholar
- Wilson MR, Sample HA, Zorn KC, Arevalo S, Yu G, Neuhaus J, et al. Clinical metagenomic sequencing for diagnosis of meningitis and encephalitis. N Engl J Med. 2019;380:2327–40.PubMedGoogle Scholar
- Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al.; China Novel Coronavirus Investigating and Research Team. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382:727–33. DOIPubMedGoogle Scholar
- Smith DJ, Gold JAW, Chiller T, Bustamante ND, Marinissen MJ, Rodriquez GG, et al.; Fungal Meningitis Response Team. Update on outbreak of fungal meningitis among U.S. residents who received epidural anesthesia at two clinics in Matamoros, Mexico. Clin Infect Dis. 2024;78:1554–8.PubMedGoogle Scholar
- US Centers for Disease Control and Prevention (CDC). Fungal meningitis outbreak associated with procedures performed under epidural anesthesia in Matamoros, Mexico [cited 2024 May 12]. https://www.cdc.gov/hai/outbreaks/meningitis-epidural-anesthesia.html
- Strong N, Meeks G, Sheth SA, McCullough L, Villalba JA, Tan C, et al. Neurovascular complications of iatrogenic Fusarium solani meningitis. N Engl J Med. 2024;390:522–9.PubMedGoogle Scholar
- Miller S, Naccache SN, Samayoa E, Messacar K, Arevalo S, Federman S, et al. Laboratory validation of a clinical metagenomic sequencing assay for pathogen detection in cerebrospinal fluid. Genome Res. 2019;29:831–42.PubMedGoogle Scholar
- Naccache SN, Federman S, Veeraraghavan N, Zaharia M, Lee D, Samayoa E, et al. A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res. 2014;24:1180–92. DOIPubMedGoogle Scholar
- Zaharia M, Bolosky B, Curtis K, Patterson D, Fox A, Patterson D, et al. Alignment in a SNAP: cancer diagnosis in the genomic age. Abstract 1912: United States and Canadian Academy of Pathology Annual Meeting; March 17–23, 2012; Vancouver, British Columbia, Canada. Lab Invest. 2012;92:458A.
- Mount DW. Using the Basic Local Alignment Search Tool (BLAST). CSH Protoc. 2007;2007:pdb.top17.
- Harrison PW, Amode MR, Austine-Orimoloye O, Azov AG, Barba M, Barnes I, et al. Ensembl 2024. Nucleic Acids Res. 2024;52(D1):D891–9.PubMedGoogle Scholar
- Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–77.PubMedGoogle Scholar
- Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80.PubMedGoogle Scholar
- Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21.PubMedGoogle Scholar
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9.PubMedGoogle Scholar
- Sayers EW, Cavanaugh M, Clark K, Pruitt KD, Sherry ST, Yankie L, et al. GenBank 2024 Update. Nucleic Acids Res. 2024;52(D1):D134–7.PubMedGoogle Scholar
- Galardini M, Mengoni A, Bazzicalupo M. Mapping contigs using CONTIGuator. Methods Mol Biol. 2015;1231:163–76.PubMedGoogle Scholar
- Lemmon AR, Brown JM, Stanger-Hall K, Lemmon EM. The effect of ambiguous data on phylogenetic estimates obtained by maximum likelihood and Bayesian inference. Syst Biol. 2009;58:130–45.PubMedGoogle Scholar
- Debourgogne A, Gueidan C, Hennequin C, Contet-Audonneau N, de Hoog S, Machouart M. Development of a new MLST scheme for differentiation of Fusarium solani Species Complex (FSSC) isolates. J Microbiol Methods. 2010;82:319–23.PubMedGoogle Scholar
- Chang DC, Grant GB, O’Donnell K, Wannemuehler KA, Noble-Wang J, Rao CY, et al.; Fusarium Keratitis Investigation Team. Multistate outbreak of Fusarium keratitis associated with use of a contact lens solution. JAMA. 2006;296:953–63. DOIPubMedGoogle Scholar
- Quick J, Grubaugh ND, Pullan ST, Claro IM, Smith AD, Gangavarapu K, et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc. 2017;12:1261–76. DOIPubMedGoogle Scholar
- Kilian M, Scholz CF, Lomholt HB. Multilocus sequence typing and phylogenetic analysis of Propionibacterium acnes. J Clin Microbiol. 2012;50:1158–65. DOIPubMedGoogle Scholar
- Yeo M, Mauricio IL, Messenger LA, Lewis MD, Llewellyn MS, Acosta N, et al. Multilocus sequence typing (MLST) for lineage assignment and high resolution diversity studies in Trypanosoma cruzi. PLoS Negl Trop Dis. 2011;5:
e1049 . DOIPubMedGoogle Scholar - O’Donnell K, Sutton DA, Fothergill A, McCarthy D, Rinaldi MG, Brandt ME, et al. Molecular phylogenetic diversity, multilocus haplotype nomenclature, and in vitro antifungal resistance within the Fusarium solani species complex. J Clin Microbiol. 2008;46:2477–90. DOIPubMedGoogle Scholar
- Tsang AKL, Lee HH, Yiu SM, Lau SKP, Woo PCY. Failure of phylogeny inferred from multilocus sequence typing to represent bacterial phylogeny. Sci Rep. 2017;7:4536. DOIPubMedGoogle Scholar
- Bleidorn C, Gerth M. A critical re-evaluation of multilocus sequence typing (MLST) efforts in Wolbachia. FEMS Microbiol Ecol. 2018;94:1. DOIPubMedGoogle Scholar
Page created: March 21, 2025
Page updated: April 25, 2025
Page reviewed: April 25, 2025
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.