Environmental and Phylogenetic Investigations of Aspergillus flavus Outbreak Linked to Contaminated Building Materials, Denmark, 2025
Alexander Gewecke, Raphael Niklaus Sieber, Søren Hallstrøm, Marc Stegger, Barbara Kolarik, Jenny D. Knudsen, Birgitte Andersen, Astrid Hall, Nadja Hawwa Vissing, and Maiken Cavling Arendrup

Author affiliation: Statens Serum Institut, Copenhagen, Denmark (A. Gewecke, R.N. Sieber, S. Hallstrøm, M. Stegger, M.C. Arendrup); Harry Butler Institute, Murdoch University, Perth, Western Australia, Australia (M. Stegger); dansk MiljøAnalyse, Vedbæk, Denmark (B. Kolarik); Copenhagen University Hospital, Rigshospitalet, Copenhagen (J.D. Knudsen, A. Hall, N.H. Vissing, M.C. Arendrup); Aalborg University, Copenhagen (B. Andersen); University of Copenhagen, Copenhagen (M.C. Arendrup)
Main Article
Figure 5
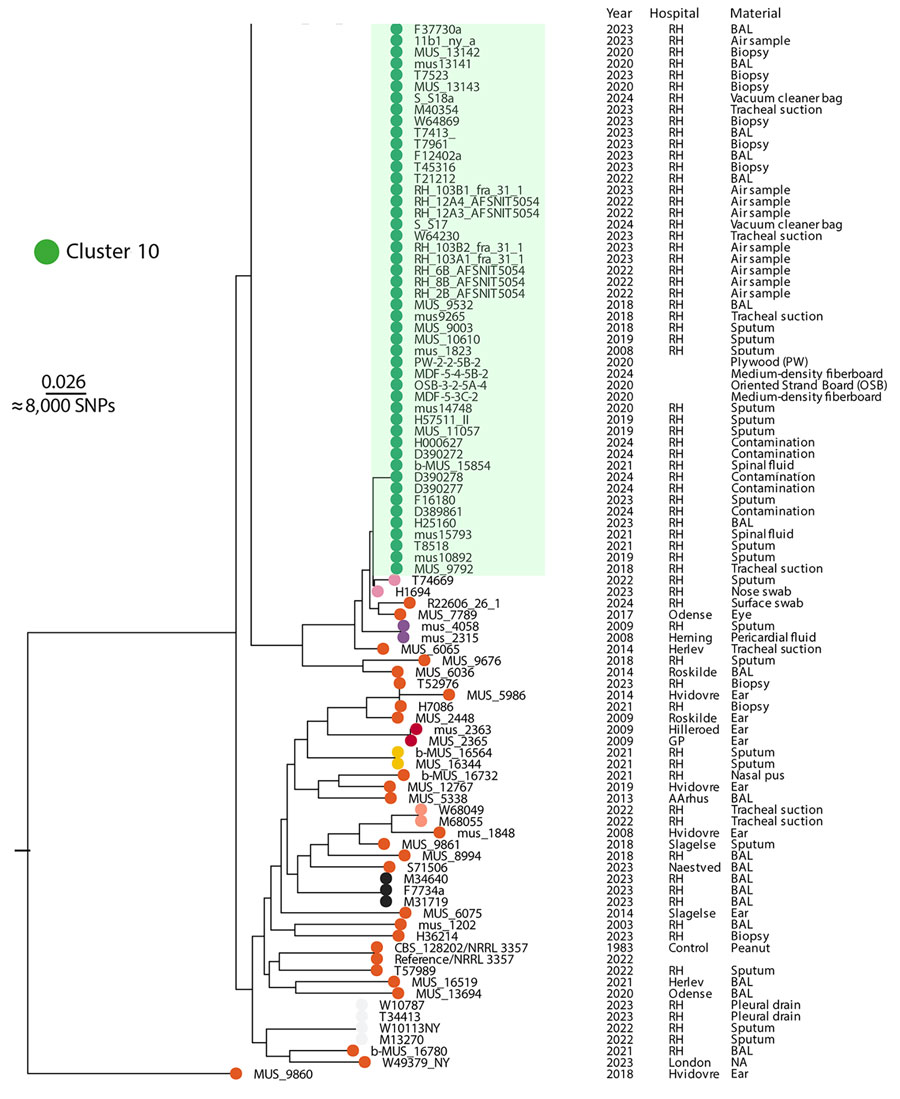
Figure 5. Rooted maximum-likelihood phylogeny of 167 A. flavus isolates (cropped) from study of environmental and phylogenetic investigations of A. flavus outbreak linked to contaminated building materials, Denmark, 2025. The tree shows clear distinction of several major clusters, including 1 monophyletic cluster 10 that contained all outbreak isolates. The tree was reconstructed based on 303,560 core-genome SNPs and rooted with MUS_9860 as outgroup. Scale bar indicates substitutions per site. Clusters were determined using TreeCluster (https://github.com/niemasd/TreeCluster). Full tree available in Appendix 1. BAL, bronchoalveolar lavage; NA, not available (information on material source); RH, Rigshospitalet; SNP, single-nucleotide polymorphism.
Main Article
Page created: January 21, 2026
Page updated: March 20, 2026
Page reviewed: March 20, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
