Volume 32, Number 3—March 2026
Research
Environmental and Phylogenetic Investigations of Aspergillus flavus Outbreak Linked to Contaminated Building Materials, Denmark, 2025
Figure 6
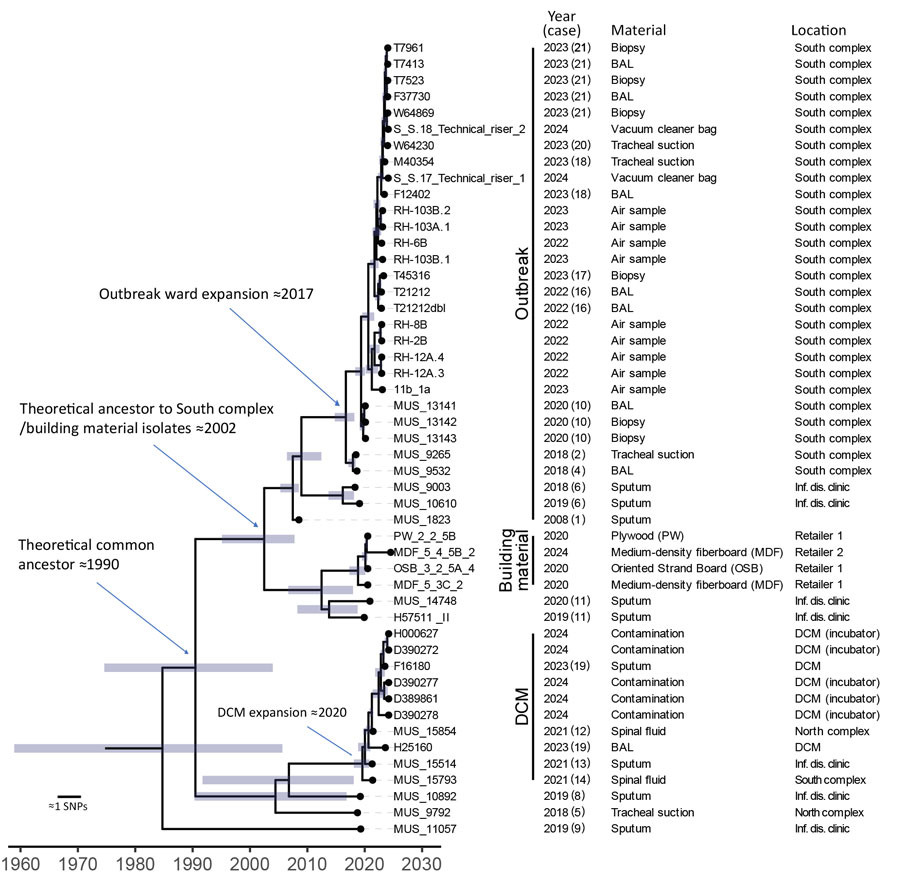
Figure 6. Time-scaled core-genome SNP-based phylogeny of cluster 10, which contained all outbreak isolates, in study of environmental and phylogenetic investigations of A. flavus outbreak linked to contaminated building materials, Denmark, 2025. Bars around internal nodes indicate the 95% highest posterior density (HPD) interval of the branching time. There is a distinct clustering of the South complex outbreak isolates (indicated as Outbreak), the isolates from building materials, and isolates from the DCM versus other quality control strains or strains from Denmark (Figure 4). Dating of ancestral nodes indicates that a common ancestor to the outbreak floor clonal expansion existed in the South complex around 2017 (95% HPD 2015–2018). The isolates from the outbreak floor are closely related with the building material isolates and share an inferred ancestor in 2002 (95% HPD 1995–2008). An expansion of a more recent ancestor occurred in the DCM around 2020 (95% HPD 2018–2021), detected as contamination in a laboratory incubator entailing an infection in >1 patient (case 19) and possibly also contamination of clinical samples (case 12, 13, 14) based on medical records. Earliest inferred ancestor to all outbreak isolates possibly existed around 1990 (95% HPD 1975–2004). The consensus tree was obtained using BEAST2 (https://www.beast2.org) on 723 core-genome SNPs called using an internal reference (T21212). A. flavus from patients 3, 7, and 15 were not available for sequencing. BAL, bronchoalveolar lavage; DCM, Department of Clinical Microbiology; inf. dis., infectious disease; SNP, single-nucleotide polymorphism.