Volume 32, Number 5—May 2026
Dispatch
Severe Respiratory Illness and Death Associated with Outbreak of Human Rhinovirus B14 among Older Adults, France, 2024
Cite This Article
Citation for Media
Abstract
We investigated an outbreak of unknown respiratory disease and 8 deaths among older adults in a long-term care facility in France. We identified human rhinovirus (HRV) by quantitative PCR and HRV-B14 by metagenomics. We obtained 5 HRV-B14 genomes that diverged from 5 publicly available genomes. Real-time metagenomics could enable rapid clinical diagnoses.
Viral respiratory infections can be particularly severe in fragile persons, especially older adults (1). Influenza virus and respiratory syncytial virus (RSV) are known to be associated with high mortality rates among persons >65 years of age (2). However, other respiratory viruses, such as human rhinovirus (HRV), also can lead to hospitalization and death (3). HRV could become a predominant viral etiology now that effective vaccines are available to prevent severe outcomes from influenza and RSV among older adults (4).
During late fall 2024, a total of 18 of 66 older adult residents of a long-term care facility (LTCF) in southeastern France developed severe respiratory illnesses. Five patients had rapid onset cardiorespiratory decompensation and died (Figure 1), which prompted notification to health authorities at the Agence Régionale de la Santé Provence-Alpes Côte d’Azur (Marseille, France). Results of quantitative PCR (qPCR) for SARS-CoV-2, influenza viruses, and RSV were negative.
Health authorities sent residue nasopharyngeal swab samples to the microbiological and virological diagnostic laboratory of the University Hospital Institute (IHU) Méditerranée Infection in Marseille for retrospective testing. We report the rapid multiplex qPCR and metagenomic investigation of that cluster of severe and fatal respiratory infections.
At IHU, we initially received 11 nasopharyngeal samples collected 3–10 days earlier from 11 of the LTCF patients. We first attempted virus isolation in a Biosafety Level 3 laboratory, as previously described (5). Within 3 hours of receipt, we tested samples by FilmArray Respiratory Panel 2 multiplex qPCR (BioFire Diagnostics, https://www.biofiredx.com); 4 samples tested HRV RNA–positive. All attempted cultures were negative.
The next day, we extracted DNA and RNA for next-generation sequencing (NGS) (Appendix). We performed qPCR on nucleic acid extracts by using the FTD Respiratory Pathogens 21 Assay (Fast Track Diagnostics/Siemens Healthineers, https://www.siemens-healthineers.com) and in-house designed qPCR (Appendix). Results of qPCR targeting 46 pathogens were negative for all but 1 sample, which was Staphylococcus aureus DNA–positive.
We then performed NGS on DNA and RNA from the 4 HRV-positive, 2 additional HRV-negative samples, and a negative, sample-free control by using the Ligation Sequencing Kit for library preparation and sequencing on GridION (Oxford Nanopore Technologies, https://nanoporetech.com). Next, we sequenced on the MiSeq platform using MiSeq Reagent version 2 kit (Illumina Inc., https://www.illumina.com), according to manufacturer’s protocol (Appendix).
We conducted bioinformatic analyses of generated NGS reads by using various commercial and freely available tools, as well as in-house pipelines (Appendix). Six hours after starting NGS, 15 NGS reads generated from the samples of 2 patients (13 reads from 1 patient and 2 reads from the other), ranging from 468 to 2,024 nt in length, were found to be best matches with an HRV-B14 genome (GenBank accession no. NC_001490.1) (Table; Appendix Tables 1, 2). We used those reads to obtain consensus sequences, one of 4,447 nt in 3 contigs for one sample, and the other of 1,019 nt for the other sample (Appendix). The negative control had no HRV reads.
We designed in-house PCR systems by using the Primer3Plus web application (https://www.primer3plus.com/index.html) to fill gaps in reconstructed HRV-B14 genomes (Appendix Table 3). Then, we used IQ-TREE (http://www.iqtree.org) to select the best model for each alignment and built maximum-likelihood phylogenetic trees. We visualized trees incorporating all complete HRV-B14 genomes by using iTOL version 7 (https://itol.embl.de).
Five days after we received the initial samples, we received additional samples from the LTCF outbreak; in total, multiplex qPCRs eventually detected HRV RNA in 13/19 nasopharyngeal swab samples from 10/13 patients. Eight samples tested positive with the BioFire assay and negative with the FTD assay (Table; Appendix Table 2). An HRV-B14–specific qPCR we designed from partial genomes was positive for 5 patients (Appendix Table 3).
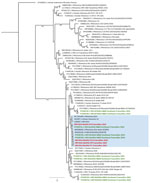
Pre-NGS viral nucleic acid enrichment using the Respiratory Viral Research Panel (Twist Bioscience, https://www.twistbioscience.com) enabled us to obtain 5 partial HRV-B14 genomes from 5 patients. Genomes were 913–6,644 nt in length and 13%–86% of the GenBank reference genome (accession no. NC_001490.1). In the phylogenetic tree, those 5 genomes clustered together with 100% bootstrap value near 5 other genomes available in public databases (Figure 2). However, 1 genome from patient P3 stood apart; its nucleotide similarity was 97.1%–98.1% with the 4 other genomes, whereas those 4 genomes had 99.9%–100.0% similarity to each other. Sequencing of the HRV virus capsid protein (VP) 1–encoding gene confirmed HRV-B14 for 4 of the initial 11 samples tested from the LTCF (Table).
We also tested 86 residue nasopharyngeal swab samples collected in southeastern France as part of routine diagnostic testing during the same timeframe as the cases from the LTC facility during late autumn 2024. Viral capsid VP1–encoding virus genotyping for all 86 nasopharyngeal swab samples identified 9 additional HRV-B sequences, 3 of which were HRV-B14 on the basis of BLAST (https://blast.ncbi.nlm.nih.gov) similarity and phylogeny (Figure 3).
The outbreak among LTCF residents resulted in 18 severe illnesses, from which 8 (44%) patients died. Of note, metagenomic methods enabled us to identify HRV-B14 within ≈30 hours of receiving samples; without those methods, the cause of the outbreak would have largely remained elusive.
Viral nucleic acid enrichment successfully lengthened the HRV-B14 genomes from the LTCF outbreak, 1 of which greatly differed from the 4 others, indicating >2 closely related HRV-B14 strains were circulating during this outbreak. In addition, HRV-B14 was detected in 3% of HRV-positive samples received at our laboratory during the same period as the LTCF outbreak, indicating that HRV-B14 circulated in that geographic area in fall of 2024.
Previous metagenomics-based approaches have successfully identified various pathogens during outbreak investigations (6–10). For instance, in the United States, metagenomics used on 6 nasopharyngeal swab samples from 4 case-patients in a pulmonary ward and on 10 nasopharyngeal swab samples from 9 control outpatients enabled investigators to determine that case-patients’ HRV sequences were not genetically related, dispelling concerns of single-source nosocomial transmission <24 hours of receiving samples (7). That case demonstrates the benefits of metagenomics in routine settings.
During the LTCF outbreak, S. aureus DNA was the only other pathogen detected by qPCR, and only in 1 sample. Notwithstanding that finding, co-infections might have been missed because specific qPCR, serologic, or culture assays were not used for all possible infectious agents; because viral or microbial loads were too low for detection; or because our laboratory was not involved in first-line diagnosis and the time between respiratory sample collection and testing at our laboratory was up to 10 days.
Severe and life-threatening HRV infections previously have been reported among older persons (11,12). Illness severity as assessed through rates of hospital admission, intensive care unit admission, and death in those studies were similar to those previously reported among immunocompromised adults infected with the 2009 influenza A(H1N1) pandemic virus (11,12) or to rates among hematopoietic cell transplant recipients with RSV or influenza virus infection (13,14). We speculate that HRV could lead to severe disease and death in fragile older persons, as seen with RSV (1), but data are scarce. Moreover, those prior assessments did not address whether severity differed between viral genotypes, and no specific data on HRV-B14 epidemiology, infectiousness, or clinical severity are available.
In summary, our findings indicate >1 HRV-B14 strain was circulating and caused severe illness and death among a population of older adults in southeastern France. Rapid identification (≈30 hours) of the viral etiology in this case demonstrates that expanded use of real-time metagenomics for routine diagnostic testing in clinical virology laboratories, particularly targeting respiratory viruses other than vaccine-preventable influenza viruses and RSV, could enable early detection and management of such respiratory pathogens.
Dr. Andreani is a virologist and junior investigator at IHU Méditerranée Infection of Aix-Marseille University and Marseille Public and University Hospitals. His research interests cover respiratory viruses and giant viruses.
Acknowledgments
This study regards data that were registered on the Health Data Access Portal of Marseille Public and University hospitals (Assistance Publique-Hôpitaux de Marseille [AP-HM]) and was approved by the Ethics and Scientific Committee of AP-HM (approval nos. CSE_PADS25-3 and CSE_PADS25-49).
Human rhinovirus B14 sequences from this study were deposited into GenBank (accession nos. PV539734–42 and PV477051–8) (Appendix Tables 1, 2).
This work was supported by the French government under the Investments for the Future program managed by the National Agency for Research, Méditerranée-Infection 10-IAHU-03, and by the Contrat Plan Etat-Région and the European funding FEDER IHUPERF. Funding sources had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; and preparation, review, or approval of the manuscript.
B.L.S. and P.C. are scientific advisors for BioSellal and Triber Global.
References
- Tseng HF, Sy LS, Ackerson B, Solano Z, Slezak J, Luo Y, et al. Severe morbidity and short- and mid- to long-term mortality in older adults hospitalized with respiratory syncytial virus infection. J Infect Dis. 2020;222:1298–310. DOIPubMedGoogle Scholar
- Godefroy R, Giraud-Gatineau A, Jimeno MT, Edouard S, Meddeb L, Zandotti C, et al. Respiratory syncytial virus infection: its propensity for bacterial coinfection and related mortality in elderly adults. Open Forum Infect Dis. 2020;7:
ofaa546 . DOIPubMedGoogle Scholar - Giraud-Gatineau A, Colson P, Jimeno MT, Zandotti C, Ninove L, Boschi C, et al. Comparison of mortality associated with respiratory viral infections between December 2019 and March 2020 with that of the previous year in Southeastern France. Int J Infect Dis. 2020;96:154–6. DOIPubMedGoogle Scholar
- Bajema KL, Yan L, Li Y, Argraves S, Rajeevan N, Fox A, et al. Respiratory syncytial virus vaccine effectiveness among US veterans, September, 2023 to March, 2024: a target trial emulation study. Lancet Infect Dis. 2025;25:625–33. DOIPubMedGoogle Scholar
- Wurtz N, Penant G, Jardot P, Duclos N, La Scola B. Culture of SARS-CoV-2 in a panel of laboratory cell lines, permissivity, and differences in growth profile. Eur J Clin Microbiol Infect Dis. 2021;40:477–84. DOIPubMedGoogle Scholar
- Slavov SN. Routine detection of viruses through metagenomics: where do we stand? Am J Trop Med Hyg. 2025;112:479–80. DOIPubMedGoogle Scholar
- Greninger AL, Waghmare A, Adler A, Qin X, Crowley JL, Englund JA, et al. Rule-out outbreak: 24-hour metagenomic next-generation sequencing for characterizing respiratory virus source for infection prevention. J Pediatric Infect Dis Soc. 2017;6:168–72. DOIPubMedGoogle Scholar
- Greninger AL, Zerr DM, Qin X, Adler AL, Sampoleo R, Kuypers JM, et al. Rapid metagenomic next-generation sequencing during an investigation of hospital-acquired human parainfluenza virus 3 infections. J Clin Microbiol. 2016;55:177–82. DOIPubMedGoogle Scholar
- Javaid W, Ehni J, Gonzalez-Reiche AS, Carreño JM, Hirsch E, Tan J, et al. Real-time investigation of a large nosocomial influenza A outbreak informed by genomic epidemiology. Clin Infect Dis. 2021;73:e4375–83. DOIPubMedGoogle Scholar
- Li P, Wang K, Qiu S, Lin Y, Xie J, Li J, et al. Rapid identification and metagenomics analysis of the adenovirus type 55 outbreak in Hubei using real-time and high-throughput sequencing platforms. Infect Genet Evol. 2021;93:
104939 . DOIPubMedGoogle Scholar - Nicholson KG, Kent J, Hammersley V, Cancio E. Risk factors for lower respiratory complications of rhinovirus infections in elderly people living in the community: prospective cohort study. BMJ. 1996;313:1119–23. DOIPubMedGoogle Scholar
- Hung IFN, Zhang AJ, To KKW, Chan JFW, Zhu SHS, Zhang R, et al. Unexpectedly higher morbidity and mortality of hospitalized elderly patients associated with rhinovirus compared with influenza virus respiratory tract infection. Int J Mol Sci. 2017;18:259. DOIPubMedGoogle Scholar
- Kraft CS, Jacob JT, Sears MH, Burd EM, Caliendo AM, Lyon GM. Severity of human rhinovirus infection in immunocompromised adults is similar to that of 2009 H1N1 influenza. J Clin Microbiol. 2012;50:1061–3. DOIPubMedGoogle Scholar
- Seo S, Waghmare A, Scott EM, Xie H, Kuypers JM, Hackman RC, et al. Human rhinovirus detection in the lower respiratory tract of hematopoietic cell transplant recipients: association with mortality. Haematologica. 2017;102:1120–30. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: May 01, 2026
Table of Contents – Volume 32, Number 5—May 2026
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Philippe Colson and Bernard La Scola, IHU Méditerranée Infection, 19-21 Blvd Jean Moulin, 13005 Marseille, France
Top