Volume 32, Number 5—May 2026
Dispatch
One Health Investigation into Fatal Encephalitis Caused by Pigeon Paramyxovirus Type 1, France
Figure 2
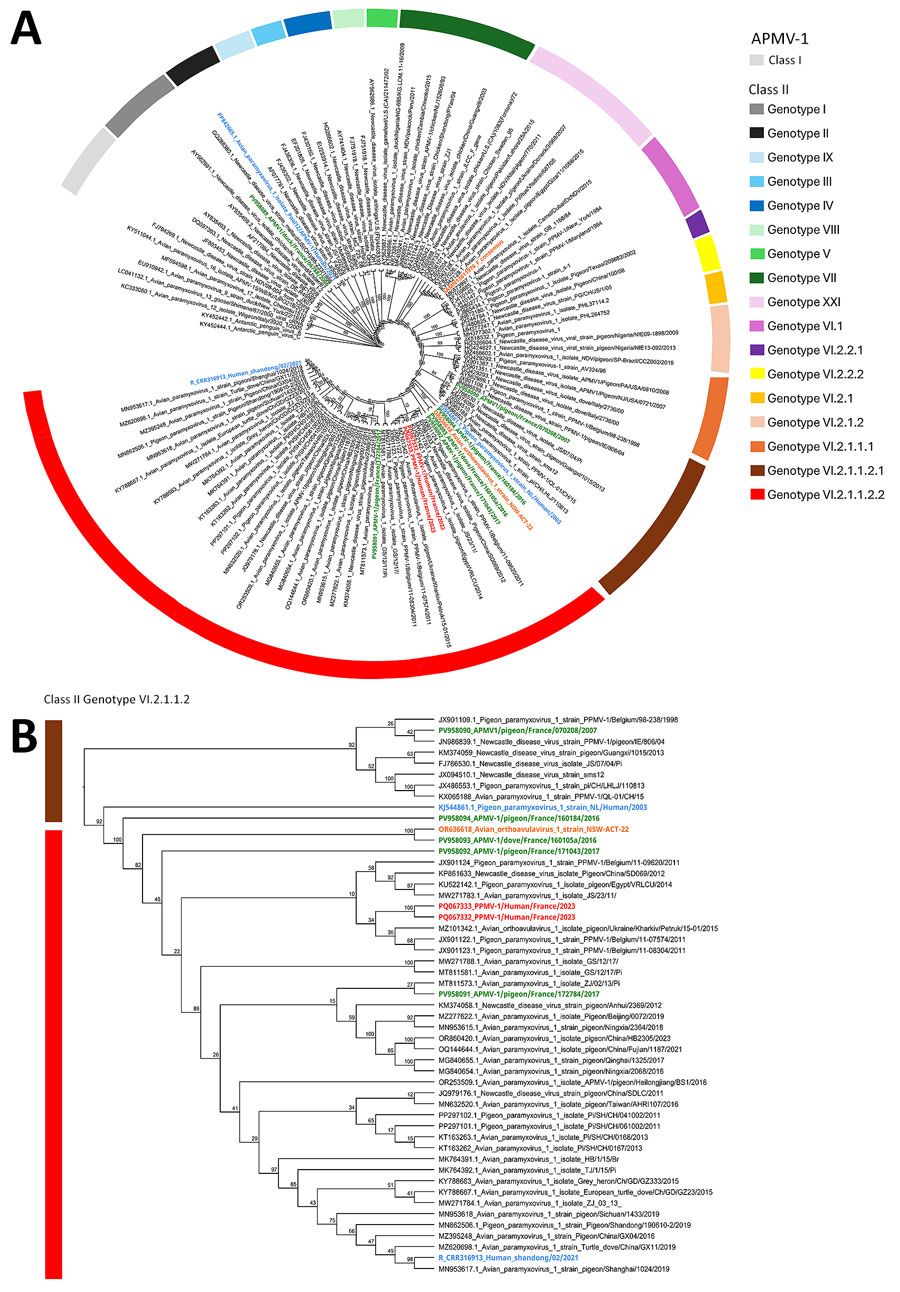
Figure 2. Phylogenetic analysis of fusion gene sequences of PPMV-1from a human case of fatal PPMV-1encephalitis in France and representative APMV-1 strains. A) We conducted phylogenetic analysis on the basis of the nucleic acid sequence of the F gene by using the maximum-likelihood method with the general time reversible with invariable site plus discrete gamma model substitution and the approximate Bayes test for the branch support analysis. Within class II, each genotype is represented by a different color. Names of sequences of interest are colored: red, PPMV-1/Human/France/2023 from the midbrain and cervical cord; green, APMV-1 strains identified in Columbidae species in France n; blue, known APMV-1 strains that caused fatal pneumonia in humans; orange, 2 strains of APMV-1 that caused encephalitis in humans. B) Class II genotype VI.2.1.1.2 PPMV-1/Human/France/2023 isolate from the patient and close reference sequences. The polybasic cleavage motif 112RRQKRF117 associated with virulent strains is exhibited by all APMV-1 class II, genotype VI, subgenotype 2.1.1.2.2 strains represented in the tree. Represented as a cladogram. Branch node values are branch support as calculated by the approximate Bayes test. GenBank accession numbers are provided. APMV-1, avian paramyxovirus type 1; PPMV-1, pigeon paramyxovirus type 1.
1These authors contributed equally to this article.
2These senior authors contributed equally to this article.