Volume 15, Number 2—February 2009
Research
Simian T-Lymphotropic Virus Diversity among Nonhuman Primates, Cameroon
Figure 3
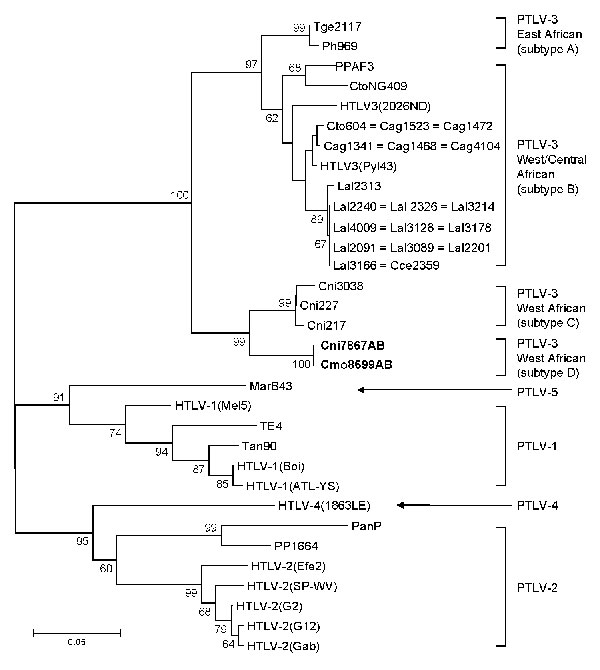
Figure 3. Identification of a novel primate T-lymphotropic virus (PTLV)–3 subtype by phylogenetic inference of 202-bp tax sequences with PTLV prototypes and partial sequences from 3 Cercopithecus nictitans (Cni217, Cni227, and Cni3038) reported elsewhere (9,21) and those identified in the current study (in boldface). GenBank accession numbers for the previously reported partial simian T-lymphotropic virus (STLV)–3 tax sequences included in this analysis are AY039033, AF412120, and AM746647–AM746673). Support for the branching order was determined by 1,000 bootstrap replicates; only values >60% are shown. Branch lengths are proportional to the evolutionary distance (scale bar) between the taxa. See Figure 2 legend for abbreviations.
References
- Calattini S, Chevalier SA, Duprez R, Bassot S, Froment A, Mahieux R, Discovery of a new human T-cell lymphotropic virus (HTLV-3) in Central Africa. Retrovirology. 2005;2:30. DOIPubMedGoogle Scholar
- Gessain A, Mahieux R. Epidemiology, origin and genetic diversity of HTLV-1 retrovirus and STLV-1 simian affiliated retrovirus [in French]. Bull Soc Pathol Exot. 2000;93:163–71.PubMedGoogle Scholar
- Mahieux R, Chappey C, Georges-Courbot MC, Dubreuil G, Mauclere P, Georges A, Simian T-cell lymphotropic virus type 1 from Mandrillus sphinx as a simian counterpart of human T-cell lymphotropic virus type 1 subtype D. J Virol. 1998;72:10316–22.PubMedGoogle Scholar
- Meertens L, Rigoulet J, Mauclere P, Van Beveren M, Chen GM, Diop O, Molecular and phylogenetic analyses of 16 novel simian T cell leukemia virus type 1 from Africa: close relationship of STLV-1 from Allenopithecus nigroviridis to HTLV-1 subtype B strains. Virology. 2001;287:275–85. DOIPubMedGoogle Scholar
- Slattery JP, Franchini G, Gessain A. Genomic evolution, patterns of global dissemination, and interspecies transmission of human and simian T-cell leukemia/lymphotropic viruses. Genome Res. 1999;9:525–40.PubMedGoogle Scholar
- Van Brussel M, Salemi M, Liu HF, Goubau P, Desmyter J, Vandamme AM. The discovery of two new divergent STLVs has implications for the evolution and epidemiology of HTLVs. Rev Med Virol. 1999;9:155–70. DOIPubMedGoogle Scholar
- Wolfe ND, Heneine W, Carr JK, Garcia AD, Shanmugam V, Tamoufe U, Emergence of unique primate T-lymphotropic viruses among central African bushmeat hunters. Proc Natl Acad Sci U S A. 2005;102:7994–9. DOIPubMedGoogle Scholar
- Van Dooren S, Meertens L, Lemey P, Gessain A, Vandamme AM. Full-genome analysis of a highly divergent simian T-cell lymphotropic virus type 1 strain in Macaca arctoides. J Gen Virol. 2005;86:1953–9. DOIPubMedGoogle Scholar
- Liégeois F, Lafay B, Switzer WM, Locatelli S, Mpoudi-Ngole E, Loul S, Identification and molecular characterization of new STLV-1 and STLV-3 strains in wild-caught nonhuman primates in Cameroon. Virology. 2008;371:405–17. DOIPubMedGoogle Scholar
- Araujo A, Hall WW. Human T-lymphotropic virus type II and neurological disease. Ann Neurol. 2004;56:10–9. DOIPubMedGoogle Scholar
- Proietti FA, Carneiro-Proietti AB, Catalan-Soares BC, Murphy EL. Global epidemiology of HTLV-I infection and associated diseases. Oncogene. 2005;24:6058–68. DOIPubMedGoogle Scholar
- Roucoux DF, Murphy EL. The epidemiology and disease outcomes of human T-lymphotropic virus type II. AIDS Rev. 2004;6:144–54.PubMedGoogle Scholar
- Yamashita M, Ido E, Miura T, Hayami M. Molecular epidemiology of HTLV-I in the world. J Acquir Immune Defic Syndr Hum Retrovirol. 1996;13(Suppl 1):S124–31. DOIPubMedGoogle Scholar
- Calattini S, Chevalier SA, Duprez R, Afonso P, Froment A, Gessain A, Human T-cell lymphotropic virus type 3: complete nucleotide sequence and characterization of the human tax3 protein. J Virol. 2006;80:9876–88. DOIPubMedGoogle Scholar
- Switzer WM, Qari SH, Wolfe ND, Burke DS, Folks TM, Heneine W. Ancient origin and molecular features of the novel human T-lymphotropic virus type 3 revealed by complete genome analysis. J Virol. 2006;80:7427–38. DOIPubMedGoogle Scholar
- Koralnik IJ, Boeri E, Saxinger WC, Monico AL, Fullen J, Gessain A, Phylogenetic associations of human and simian T-cell leukemia/lymphotropic virus type I strains: evidence for interspecies transmission. J Virol. 1994;68:2693–707.PubMedGoogle Scholar
- Meertens L, Gessain A. Divergent simian T-cell lymphotropic virus type 3 (STLV-3) in wild-caught Papio hamadryas papio from Senegal: widespread distribution of STLV-3 in Africa. J Virol. 2003;77:782–9. DOIPubMedGoogle Scholar
- Goubau P, Van Brussel M, Vandamme AM, Liu HF, Desmyter J. A primate T-lymphotropic virus, PTLV-L, different from human T-lymphotropic viruses types I and II, in a wild-caught baboon (Papio hamadryas). Proc Natl Acad Sci U S A. 1994;91:2848–52. DOIPubMedGoogle Scholar
- Van Dooren S, Shanmugam V, Bhullar V, Parekh B, Vandamme AM, Heneine W, Identification in gelada baboons (Theropithecus gelada) of a distinct simian T-cell lymphotropic virus type 3 with a broad range of Western blot reactivity. J Gen Virol. 2004;85:507–19. DOIPubMedGoogle Scholar
- Meertens L, Mahieux R, Mauclere P, Lewis J, Gessain A. Complete sequence of a novel highly divergent simian T-cell lymphotropic virus from wild-caught red-capped mangabeys (Cercocebus torquatus) from Cameroon: a new primate T-lymphotropic virus type 3 subtype. J Virol. 2002;76:259–68. DOIPubMedGoogle Scholar
- Van Dooren S, Salemi M, Pourrut X, Peeters M, Delaporte E, Van Ranst M, Evidence for a second simian T-cell lymphotropic virus type 3 in Cercopithecus nictitans from Cameroon. J Virol. 2001;75:11939–41. DOIPubMedGoogle Scholar
- Song KJ, Nerurkar VR, Saitou N, Lazo A, Blakeslee JR, Miyoshi I, Genetic analysis and molecular phylogeny of simian T-cell lymphotropic virus type I: evidence for independent virus evolution in Asia and Africa. Virology. 1994;199:56–66. DOIPubMedGoogle Scholar
- Van Dooren S, Verschoor EJ, Fagrouch Z, Vandamme AM. Phylogeny of primate T lymphotropic virus type 1 (PTLV-1) including various new Asian and African non-human primate strains. Infect Genet Evol. 2007;7:374–81. DOIPubMedGoogle Scholar
- Kingdon J. The Kingdon field guide to African mammals. London: Academic Press; 1997.
- Courgnaud V, Van Dooren S, Liegeois F, Pourrut X, Abela B, Loul S, Simian T-cell leukemia virus (STLV) infection in wild primate populations in Cameroon: evidence for dual STLV type 1 and type 3 infection in agile mangabeys (Cercocebus agilis). J Virol. 2004;78:4700–9. DOIPubMedGoogle Scholar
- Switzer WM, Salemi M, Shanmugam V, Gao F, Cong ME, Kuiken C, Ancient co-speciation of simian foamy viruses and primates. Nature. 2005;434:376–80. DOIPubMedGoogle Scholar
- Busch MP, Switzer WM, Murphy EL, Thomson R, Heneine W. Absence of evidence of infection with divergent primate T-lymphotropic viruses in United States blood donors who have seroindeterminate HTLV test results. Transfusion. 2000;40:443–9. DOIPubMedGoogle Scholar
- Wolfe ND, Switzer WM, Carr JK, Bhullar VB, Shanmugam V, Tamoufe U, Naturally acquired simian retrovirus infections in central African hunters. Lancet. 2004;363:932–7. DOIPubMedGoogle Scholar
- Lemey P, Pybus OG, Van Dooren S, Vandamme AM. A Bayesian statistical analysis of human T-cell lymphotropic virus evolutionary rates. Infect Genet Evol. 2005;5:291–8. DOIPubMedGoogle Scholar
- Salemi M, Desmyter J, Vandamme AM. Tempo and mode of human and simian T-lymphotropic virus (HTLV/STLV) evolution revealed by analyses of full-genome sequences. Mol Biol Evol. 2000;17:374–86.PubMedGoogle Scholar
- Meertens L, Shanmugam V, Gessain A, Beer BE, Tooze Z, Heneine W, A novel, divergent simian T-cell lymphotropic virus type 3 in a wild-caught red-capped mangabey (Cercocebus torquatus torquatus) from Nigeria. J Gen Virol. 2003;84:2723–7. DOIPubMedGoogle Scholar
- Hahn BH, Shaw GM, De Cock KM, Sharp PM. AIDS as a zoonosis: scientific and public health implications. Science. 2000;287:607–14. DOIPubMedGoogle Scholar
- Salemi M, Van Dooren S, Audenaert E, Delaporte E, Goubau P, Desmyter J, Two new human T-lymphotropic virus type I phylogenetic subtypes in seroindeterminates, a Mbuti pygmy and a Gabonese, have closest relatives among African STLV-I strains. Virology. 1998;246:277–87. DOIPubMedGoogle Scholar
- Aghokeng AF, Liu W, Bibollet-Ruche F, Loul S, Mpoudi-Ngole E, Laurent C, Widely varying SIV prevalence rates in naturally infected primate species from Cameroon. Virology. 2006;345:174–89. DOIPubMedGoogle Scholar
- Peeters M, Courgnaud V, Abela B, Auzel P, Pourrut X, Bibollet-Ruche F, Risk to human health from a plethora of simian immunodeficiency viruses in primate bushmeat. Emerg Infect Dis. 2002;8:451–7.PubMedGoogle Scholar
- Wolfe ND, Dunavan CP, Diamond J. Origins of major human infectious diseases. Nature. 2007;447:279–83. DOIPubMedGoogle Scholar
- Wolfe ND, Switzer WM, Heneine W. Emergence of novel retroviruses. Washington: American Society for Microbiology Press; 2006.
1Current affiliation: Stanford University, Stanford, California, USA.
2Current affiliation: University of Pittsburgh Graduate School of Public Health, Pittsburgh, Pennsylvania, USA.