Volume 16, Number 11—November 2010
Research
Genetic Structure of Plasmodium falciparum and Elimination of Malaria, Comoros Archipelago
Cite This Article
Citation for Media
Abstract
The efficacy of malaria control and elimination on islands may depend on the intensity of new parasite inflow. On the Comoros archipelago, where falciparum malaria remains a major public health problem because of spread of drug resistance and insufficient malaria control, recent interventions for malaria elimination were planned on Moheli, 1 of 4 islands in the Comoros archipelago. To assess the relevance of such a local strategy, we performed a population genetics analysis by using multilocus microsatellite and resistance genotyping of Plasmodium falciparum sampled from each island of the archipelago. We found a contrasted population genetic structure explained by geographic isolation, human migration, malaria transmission, and drug selective pressure. Our findings suggest that malaria elimination interventions should be implemented simultaneously on the entire archipelago rather than restricted to 1 island and demonstrate the necessity for specific chemoresistance surveillance on each of the 4 Comorian islands.
Plasmodium falciparum causes malaria worldwide; 250 million cases and ≈1 million deaths occur annually, mostly in sub-Saharan Africa. However, recently increased international financial commitment has revived hope for malaria elimination from selected areas to which it is endemic, and the feasibility of elimination has become a topic of research (1). The successful elimination of malaria from several Caribbean islands, Cyprus, Reunion, Mauritius, Maldives, Taiwan, and Singapore in the context of the Global Malaria Eradication Program (1955–1968) (2) suggests that islands are prime targets for elimination interventions. Because most parasites among neighboring areas are exchanged through human migrations, the geographic isolation of islands can limit malaria importation and may make control easier (3,4).
Several molecular epidemiologic studies have shown that P. falciparum populations are more or less homogeneous within malaria-endemic areas and may exhibit genetic structure patterns shaped by various transmission rates and geographic isolation levels (4–6). Although geographic isolation may be more relevant on islands than within continents, the role of parasite genetic structure in malaria-endemic archipelagos or among malaria-endemic islands and the nearest continent remains unknown. Past failures of malaria elimination in Zanzibar off the coast of mainland Tanzania; in Sri Lanka (1); or in Mayotte, a France-administered island of the Comoros archipelago (7), raise the question of the minimal geographic isolation level and the optimal size of intervention area required for malaria elimination success. Analysis of malaria epidemiology in Comoros archipelago, where a limited malaria elimination program is ongoing, may help to answer this question.
Falciparum malaria remains a major public health problem on the 4 islands of the Comoros archipelago (Grande Comore, Moheli, Anjouan, and Mayotte) (Table 1) in the Indian Ocean between Madagascar and the eastern coast of Africa. Malaria control has been hampered by the emergence of P. falciparum resistance to chloroquine and to pyrimethamine/sulfadoxine in the early 1980s (7,10) and of Anopheles mosquitoes resistance to DDT. Malaria control also has had recurrent political, economic, and structural weaknesses in the Union of the Comoros (the state comprising Grande Comore, Moheli, and Anjouan islands). Under stable political and economic conditions, notable efforts in case management and vector control in Mayotte failed to eliminate falciparum malaria and to prevent recurrent epidemics (Table 1). During the past 6 years Since 2004, health authorities in Grande Comore and France have introduced an artemisinin-based combined therapy (artemether plus lumefantrin) as first-line treatment for uncomplicated falciparum malaria (7,10). Large-scale distribution of insecticide-treated mosquito nets also has been gradually implemented on Grande Comore, Moheli, and Anjouan (8), with the goal of reaching up to 89.1% and 46.3% of the households with at least 1 mosquito net and 1 insecticide-impregnated mosquito net, respectively, among 1,620 households from the 3 islands (Comoran National Malaria Control Program, unpub. data, 2007).
In Mayotte, anti–Anopheles spp. mosquito larvae measures have been strengthened. Finally, by late 2007, a controversial malaria elimination project was launched on the sole island of Moheli with assistance from China. Mass treatment of the residing and disembarking population with artemisinin plus piperaquine (Artequick; Artepharm Co., Guangzhou, People’s Republic of China) and primaquine was initiated without enhancement of vector control. Because of continual human travel across the archipelago, the long-term success of such a spatially limited elimination attempt is questionable.
In addition, surveillance of P. falciparum chemosusceptibility has been chaotic and unequal among the islands of the archipelago, and results of the few available therapeutic efficacy tests and in vitro and molecular resistance studies often have been discordant. A more rational and efficient surveillance system is urgently needed. Because Marseille, France, houses a Comorian community of 50,000–80,000 persons who annually import several hundred malaria cases, the city was proposed as a relevant surveillance site for chemosusceptibility of P. falciparum imported from Comoros (11). However, extrapolating these chemoresistance data to the entire archipelago remains difficult.
As already proposed for Borneo (12) and the Philippines (13), our main objective was to analyze the genetic structure of P. falciparum on the Comoros islands to 1) forecast the chances of middle-term and long-term success for the current elimination program focalized in Moheli, 2) guide future malaria elimination programs on the archipelago, and 3) adjust its chemoresistance monitoring and treatment policies. Study results also would provide a pertinent model for determining which other malaria-endemic areas might be eligible for malaria elimination. A secondary objective was to assess whether the diversity of the P. falciparum strains imported into Marseille were representative of the P. falciparum populations from Comoros so we could evaluate the relevance of distant chemoresistance surveillance from Marseille.
We characterized P. falciparum populations from each of the 4 islands and from Marseille (imported from the archipelago) by multilocus microsatellite genotyping. The genetic polymorphism of 3 genes involved in P. falciparum resistance to chloroquine, pyrimethamine and cycloguanil, or sulfadoxine was also investigated.
P. falciparum Isolates
The study was conducted in 2007 (before the malaria elimination program was launched in Moheli) in each of the 4 islands of the Comoros archipelago and in Marseille. The protocol was approved by the ethics committee of the university hospitals of Marseille and by the Comorian Ministry of Health. Blood samples were obtained after informed consent from patients seeking care for symptomatic falciparum malaria at healthcare centers of the archipelago or at emergency departments of hospitals in Marseille.
Blood samples were absorbed onto Whatman FTA Elute absorbent filter paper in Grande Comore, Moheli, and Anjouan islands, on Whatman 903 Protein Saver filter paper (Whatman Inc., Florham Park, NJ, USA) in Mayotte, and collected into Vacutainer tubes (Becton Dickinson, Le Pont-De-Claix, France) in Marseille. All samples were frozen and kept at –20°C. After eliminating samples with missing data or the lowest parasitaemia levels (<0.01%), 36 isolates per site were randomly chosen for genotyping, a sample size considered adequate for the planned population genetics analyses.
Collection of Epidemiologic Data
Patient's age, sex, history of travel across or outside the archipelago (during the past year for Grande Comore, Moheli, and Anjouan; during the past 3 weeks for Mayotte) and history of recent clinical malaria episodes and intake of antimalarial drugs (during the previous month) were collected by oral questioning. Distances between each island were measured by using Google Earth software.
Genotyping Procedures
DNA was extracted from filter papers according to the manufacturer's recommendations (Whatman Inc.) and from whole blood from Vacutainer tubes by using the EZNA Blood DNA Kit (Biofidal, Vaulx-en-Velin, France). Next, the entire genome was amplified by using the Illustra GenomiPhi V2 DNA Amplification Kit (GE Healthcare, Little Chalfont, UK).
Molecular Markers
Length polymorphism was analyzed for 6 complex and putatively neutral microsatellite loci previously described (4): Pf2689, 7A11, C4M79, Pf2802, TRAP, and C4M69 (Appendix Table). The studied chemoresistance markers were the K76 point mutation of the P. falciparum chloroquine resistance transporter (Pfcrt) gene (associated with P. falciparum resistance to chloroquine) (14); point mutations of P. falciparum dihydrofolate reductase (Pfdhfr) gene codons 108, 16, 51, 59, and 164 (associated with P. falciparum resistance to pyrimethamine and cycloguanil, i.e., proguanil metabolite) (15); and P. falciparum dihydropteroate synthase (Pfdhps) gene codons 436, 437, 540, 581, and 613 (associated with P. falciparum resistance to sulfadoxine) (15) (Appendix Table).
Genotyping by PCR
Microsatellite loci were amplified by nested PCR with fluorescent end-labeled primers before electrophoresis on polyacrylamide gels with Genescan-500 LIZ labeled size standards on an ABI 3130XL capillary sequencer (Applied Biosystems, Warrington, UK) (Appendix Table). Their length was then analyzed by using GENESCAN software (Applied Biosystems, Carlsbad, CA, USA), as described (4). The Pfcrt gene was amplified by seminested PCR, and the codon 76 mutation was genotyped by using a simple PCR-restriction fragment digest assay and fluorescent detection of products on an ABI 3130XL capillary sequencer, as described (16). The Pfdhps and Pfdhfr genes were amplified by nested PCR, and their mutations were genotyped by using a primer extension method, as described (17) and electrophoresis on the ABI 3130XL capillary sequencer.
Statistical Analysis
The multiplicity of infection (MOI, i.e., the number of parasites genetically distinguishable by different alleles) with P. falciparum was estimated for each isolate from the microsatellite locus that exhibited the highest number of alleles. The mean MOI for each P. falciparum population (Grande Comore, Moheli, Anjouan, Mayotte, and Marseille) was then calculated. Each pair of sites was compared for MOI by using the Mann-Whitney U test.
For parasites with multiple infection, i.e., >1 allele at each locus, we conducted separate subsequent analysis considering the following: 1) complete dataset, 2) curtailed dataset with single or main clones after elimination of isolates unsuccessfully genotyped at all 6 microsatellite loci, or 3) reconstructed multilocus genotypes after elimination of samples with impossible reconstruction (>1 allele at >1 locus with equivocal peak intensities) and elimination of unsuccessfully genotyped isolates (Table 1).
Genetic Diversity
Genetic diversity of the 5 P. falciparum populations was assessed by the number of alleles per locus and by the Nei unbiased expected heterozygosity index (He) calculated from allelic frequencies on the 6 microsatellites for complete datasets by using GENETIX software version 4.05 (18,19). Comparison between He of the 5 different populations was performed on FSTAT software version 2.9.4 with a 1,000 permutations bilateral comparison test (20).
Population Genetic Structure
Population genetic structure was investigated by using the Wright F statistic (FST) (21). The FST index was computed for the 6 microsatellite markers and 5 populations on FSTAT software version 2.9.4 (20,22) and by using the Slatkin index on ARLEQUIN software (23). A canonical correspondence analysis of the reconstructed multilocus genotypes set was conducted to illustrate measures of population structure (24) by using CANOCO software (25), and its graphic representation was performed by using R software. A Monte Carlo procedure permuting genotypes among the populations was used to test the significance of the canonical axes and estimate the 95% confidence intervals of the centroid of each population (25).
Frequency of Mutations Associated with Chemoresistance
We estimated the frequency of point mutations on the Pfcrt (K76T mutation), Pfdhfr (108 + 59 + 51 triple mutation), and Pfdhps (437 + 540 double mutation) genes. Differences among sites were tested by using the Fisher exact test.
Associations between FST and Estimations of Parasite Flux
The association between genetic distance (transformed as FST/[1 – FST]) and the natural log of the geographic distance in kilometers was investigated for each pair of islands according to the isolation-by-distance model (5,12,26). When we considered the number of patients in each sampled island (A) with history of recent arrival from each of the neighboring islands (B) and thus possibly with imported malaria (Table 1), the relationship between FST and the mean proportion of these travelers among patients, calculated as ([NB→A/NA] + [NA→B/NB])/2, was investigated for each pair of islands.
P. falciparum was detected by PCR in each of the 36 genotyped blood samples from all 5 sites. Microsatellite genotyping was complete for 149 (83%) of the 180 samples (Table 2).
Mean MOIs
Of the 180 samples, 59 isolates were multi-infected; the proportion of multi-infection among islands differed substantially (Table 2). The mean MOI ranged from 1.22 in Mayotte to 2.11 in Anjouan (Table 3). It was significantly higher in Anjouan than in Grande Comore (p = 0.0015), Moheli (p = 0.0051), and Mayotte (p = 0.0001) and higher in Marseille than in Mayotte (p = 0.0093).
Genetic Diversity
Genetic diversity (He) of each population, estimated by unbiased expected heterozygosity based on allelic frequencies of the 6 microsatellites and the complete dataset, is shown in Table 4. The highest diversity was observed for Anjouan and Moheli (each He = 0.71) and the lowest diversity for Mayotte (He = 0.63). The mean He was significantly lower for Mayotte than for Marseille (p = 0.04) and lower than for the other sites combined (p = 0.001).
Genetic Differentiation among Islands and Population Structure
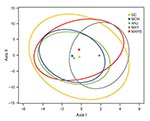
Figure 1 shows the centroid of each falciparum population surrounded by its 95% confidence interval, and both axes were significant (p = 0.0001 and p = 0.0004 for 1,000 permutations, respectively). Grande Comore, Moheli, and Anjouan nearby centroids suggest closely related populations. The detached Mayotte centroid suggests a marked differentiation from all the other populations.
Figure 2 shows the pairwise differentiation coefficients (FST) estimated for the 5 parasite populations according to the 6 microsatellite loci and the complete dataset (n = 281). The number of P. falciparum clones used to calculate FST between sites was 50, 53, 76, 44, and 58 in Grande Comore, Moheli, Anjouan, Mayotte, and Marseille, respectively. The Moheli parasite population did not differ significantly from the Grande Comore and Anjouan populations. Conversely, the Mayotte population differed significantly from the populations of the 4 other sites. Marseille parasite populations differed significantly from those from all sites except Grande Comore. Similar differentiation index were obtained by using a curtailed dataset (n = 149) or reconstructed multilocus genotypes (n = 170) and by using the Slatkin index (data not shown).
Relations between FST and Estimations of Parasite Flux
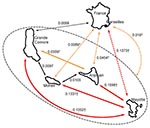
Association between genetic and geographic distances for each pair of islands is shown in Figure 3. No association was significant. The Anjouan–Mayotte pair exhibited a large FST despite the close proximity of the 2 islands.
Of the 414 patients sampled in the archipelago, 35 reported recent travel to neighboring islands (Table 1). Figure 3 suggests a negative relationship between the mean percentage of travelers among patients and the corresponding FST.
Frequency of Point Mutations associated with Chemoresistance
Prevalence of Pfcrt, Pfdhfr, and Pfdhps mutations in the 5 P. falciparum study populations are presented in Table 5. Prevalence of the Pfcrt mutation (i.e., isolates with the 76T allele or with the 76 K, and T alleles) was significantly lower in Anjouan than in the other parasite populations (p<0.002). Prevalence of the Pfcrt mutation was significantly higher in Mayotte than in any other population (p< 0.0001). The prevalences of the Pfcrt mutation in Grande Comore, Moheli, and Marseille did not differ significantly.
When mutated, the Pfdhfr gene frequently exhibited the association of the 3 mutations 108N + 59R + 51I. Prevalence of this triple mutation was significantly lower in the Anjouan population than in the Grande Comore (p = 0.04), Moheli (p = 0.003), or Marseille populations (p = 0.0004). Its prevalence also was significantly higher in Marseille than in Mayotte (p = 0.02). The prevalence of Pfdhps gene mutations appeared low in the 5 populations.
Multi-infected isolates with genotype ambiguities and impossible distinction between associated clones were rare. However, prevalence of these mutations varied little, regardless whether these ambiguous multi-infected isolates were considered.
The mean MOIs remained low for the Comoros archipelago in comparison with African areas, where malaria is highly endemic (4), most likely because of moderate levels of malaria transmission (4,5). Likewise, the significantly higher MOI in Anjouan may reflect a higher level of malaria transmission in the rainy and swampy sampled areas, where vector control has for a long time been impaired by recurrent island-specific political crises. The genetic diversities appeared lower on the archipelago than on most of the African continent (4,5,27–30), probably because of the geographic isolation of the islands and their lower malaria transmission levels that could limit effective parasite population sizes and outbreeding. However, genetic diversities remained higher than in Asia (5,6,12,13) and South America (5,31).
The genetic differentiation index (FST) exhibited a contrasted genetic structure between the studied P. falciparum populations. Genetic distances were low among parasites on Grande Comore, Moheli, and Anjouan islands. However, FSTs among these 3 populations and Mayotte were as significant as between P. falciparum populations of Senegal and Djibouti when the same microsatellite loci were used (4) or as between populations of Africa and Southeast Asia when other microsatellite loci were used (5). In addition, the genetic distances between falciparum populations across the archipelago seemed associated with the parasite flows among islands, estimated from the proportion of travelers among sampled patients, in particular for Moheli, Grande Comore, and Anjouan.
Our results strongly suggest that despite the insular geographic isolation of Moheli and the malaria elimination program launched in late 2007 on this island only, the mass treatment without enhanced vector control may soon be impaired by the continuous importation of new parasites through intense human migrations. In addition to flights and ferries regularly traveling across the archipelago, humans in Comoros move mainly by small fishing boats, especially from and toward Moheli (S. Rebaudet, pers. comm.). Their ubiquitous and informal traffic makes human flux estimations unreliable (probably several tens of thousands of persons each way annually [S. Rebaudet, pers. comm.]) and their control difficult. These factors might explain why, despite the substantial resources that France has allocated in Mayotte to malaria control since the mid-1970s, malaria importation to this island could not be stopped and autochthonous falciparum malaria could not be eliminated. The disease persists in Mayotte with a hypoendemo-epidemic setting, genetically characterized by low MOI, low He, significant linkage disequilibrium (data not shown), and high FSTs, artificially overestimated by the sampling of multiple repeated genotypes (data not shown).
The persistent efforts for malaria elimination in Moheli can be hypothesized to create a Mayotte-like setting requiring efficient vector control to prevent epidemics in a Mohelian population that is losing its immunity. In the Union of the Comoros, the extension of the elimination program based on artemisinin-based combined therapy mass treatment to Grande Comore and Anjouan is being considered by Comorian health authorities and their Chinese interlocutors (R. Silaï, pers. comm.). Its success and the prevention of epidemics will depend on the rapid and large implementation of the preventive, diagnostic, and therapeutic measures planned with the funds granted in 2010 by Round 8 of the Global Fund (http://portfolio.theglobalfund.org/Grant/Index/COM-810-G03-M?lang=en).
Isolation of a specific P. falciparum population before planning its elimination needs to be appropriately evaluated. Results from the present Comorian epidemiologic study illustrate how it could be evaluated by a population genetics approach. In that type of geographic setting, population genetics studies provide a probably more direct and reliable estimation of parasite flows and risk for re-introduction than does the evaluation of human population movements by sociodemographic methods. Therefore, the relevance of parasite inflow from Africa (mostly the Tanzania coast, Madagascar, or other malaria-endemic areas) should be evaluated before the elimination project is extended to the rest of the Comoros archipelago. Similar data would also be useful for Sri Lanka, Malaysia, Indonesia, the Philippines, Solomon islands, or Vanuatu, several islands where national or localized malaria elimination projects are being implemented (2).
According to the genetic structure of P. falciparum populations in Comoros demonstrated by microsatellite genotyping, resistance levels would be expected to be fairly similar across the archipelago, except for Mayotte. However, the study of Pfcrt and Pfdhfr resistance–associated mutations differed markedly, explainable only by contrasting levels of drug selective pressure among islands. Indeed, the prevalence of the K76T mutation on the Pfcrt gene was high in both Grande Comore and Moheli as found in previous studies (32,33) but substantively lower in Anjouan and significantly higher in Mayotte where chloroquine use was massive during 1975–2007 (7,34). Similarly, the prevalence of Pfdhfr triple mutants was higher in Moheli than in Anjouan and the prevalence of Pfdhfr double or triple mutants higher in Marseille than in Grande Comore.
Although no reliable estimation of past use of antimalarial drugs in Comoros is available, these differences may be explained by a greater use in Moheli of pyrimethamine (in the sulfadoxine/pyrimethamine combination for malaria treatment) and trimethoprim (in cotrimoxazole compound, which is widely prescribed in this island as an antimicrobial drug) and in Marseille of proguanil (in association with chloroquine or atovaquone, used as malaria chemoprophylaxis by travelers to the archipelago) (S. Rebaudet, pers. comm.). Trimethoprim and proguanil are 2 antifolate drugs whose cross-resistance with pyrimethamine has been suspected (35,36) and that may have selected these Pfdhfr mutations. Because of the contrasting resistance levels among islands, the risk for rapid propagation of resistant P. falciparum strains across the archipelago suggested by the low FSTs among Grande Comore, Moheli, and Anjouan (4,5), and the easier selection of multigenic resistance and multiresistance from low MOIs limiting the possibilities of genetic recombinations that could break apart allele combinations (5,37,38), French and Comorian health authorities should organize surveillance of chemoresistance, both regular and separated for each island.
Finally, microsatellite genotypes of the P. falciparum population in Marseille substantially differed from those populations on all islands except Grande Comore. Because most of the Comorian inhabitants living in Marseille originated from Grande Comore, malaria is imported mainly from this particular island (S. Rebaudet, pers. comm.). Therefore, if we consider that the P. falciparum population in Marseille may be representative only of the Grande Comore population and the distinct levels of drug pressure between Marseille and the other populations, the relevance of distant chemosusceptibility surveillance from Marseille is likely to be limited.
Dr Rebaudet works in the Infectious and Tropical Diseases Department of Paul-Brousse Hospital, Assistance Publique–Hôpitaux de Paris, and in the Institut de Recherche Biomédicale des Armées, Marseille. His primary research interests include tropical medicine, particularly malaria and its epidemiology on the Comoros archipelago.
Acknowledgments
We are indebted to the residents of the studied sites for their cooperation during the survey. We thank the laboratory technicians of the Institut de Recherche Biomédicale des Armées (Marseille, France) for their technical support and the staff of the Programme National de Lutte contre le Paludisme (Moroni, Union of the Comoros) and of the Direction des Affaires Sanitaires et Sociales of Mayotte for providing determinant data on malaria epidemiology. We also thank François Renaud and Frank Prugnolle for reviewing the manuscript
This work was supported by the Programme Hospitalier de Recherche Clinique Régional Assistance Publique–Hôpitaux de Marseille 2003, the Conseil Général des Bouches-du-Rhône, the French Ministry of Research, and the French Ministry of Defense (Dynapop Program, 03co007-05 from the Délégation Générale pour l’Armement; Schema Directeur de Lutte contre le Paludisme de l'Etat Major des Armées).
References
- Roll Back Malaria Partnership. The global malaria action plan, for a malaria-free world. Geneva: World Health Organization; 2008 [cited 2010 Sep 5]. http://www.rollbackmalaria.org/gmap/index.html
- World Health Organization. World malaria report 2009. Geneva: The Organization; 2009 [cited 2010 Sep 5]. http://www.who.int/malaria/publications/atoz/9789241563901/en/index.html
- Rogier C, Pradines B, Bogreau H, Koeck JL, Kamil MA, Mercereau-Puijalon O. Malaria epidemic and drug resistance, Djibouti. Emerg Infect Dis. 2005;11:317–21.PubMedGoogle Scholar
- Bogreau H, Renaud F, Bouchiba H, Durand P, Assi SB, Henry MC, Genetic diversity and structure of African Plasmodium falciparum populations in urban and rural areas. Am J Trop Med Hyg. 2006;74:953–9.PubMedGoogle Scholar
- Anderson TJ, Haubold B, Williams JT, Estrada-Franco JG, Richardson L, Mollinedo R, Microsatellite markers reveal a spectrum of population structures in the malaria parasite Plasmodium falciparum. Mol Biol Evol. 2000;17:1467–82.PubMedGoogle Scholar
- Pumpaibool T, Arnathau C, Durand P, Kanchanakhan N, Siripoon N, Suegorn A, Genetic diversity and population structure of Plasmodium falciparum in Thailand, a low transmission country. Malar J. 2009;8:155. DOIPubMedGoogle Scholar
- Quatresous I, Pettinelli F, Le Bras J, Solet JL, Lepère JF, Giry C, What do we know about malaria resistance in Mayotte, France, in 2007? [in French]. Bulletin Epidémiologique Hebdomadaire. 2007;48–49:409–12.
- World Health Organization. World malaria report 2008. Geneva: The Organization; 2008 [cited 2010 Sep 5]. http://malaria.who.int/wmr2008/malaria2008.pdf
- World Health Organization. Malaria control and elimination: report of a technical review. Geneva: The Organization; 2008 [cited 2010 Sep 5]. http://www.who.int/malaria/publications/atoz/9789241596756/en/index.html
- Silai R, Moussa M, Abdalli Mari M, Astafieva-Djaza M, Hafidhou M, Oumadi A, Surveillance of falciparum malaria susceptibility to antimalarial drugs and policy change in the Comoros [in French]. Bull Soc Pathol Exot. 2007;100:6–9. DOIPubMedGoogle Scholar
- Parola P, Pradines B, Simon F, Carlotti MP, Minodier P, Ranjeva MP, Antimalarial drug susceptibility and point mutations associated with drug resistance in 248 Plasmodium falciparum isolates imported from Comoros to Marseille, France, 2004–2006. Am J Trop Med Hyg. 2007;77:431–7.PubMedGoogle Scholar
- Anthony TG, Conway DJ, Cox-Singh J, Matusop A, Ratnam S, Shamsul S, Fragmented population structure of Plasmodium falciparum in a region of declining endemicity. J Infect Dis. 2005;191:1558–64. DOIPubMedGoogle Scholar
- Iwagami M, Rivera PT, Villacorte EA, Escueta AD, Hatabu T, Kawazu S, Genetic diversity and population structure of Plasmodium falciparum in the Philippines. Malar J. 2009;8:96. DOIPubMedGoogle Scholar
- Djimde A, Doumbo OK, Cortese JF, Kayentao K, Doumbo S, Diourte Y, A molecular marker for chloroquine-resistant falciparum malaria. N Engl J Med. 2001;344:257–63. DOIPubMedGoogle Scholar
- Kublin JG, Dzinjalamala FK, Kamwendo DD, Malkin EM, Cortese JF, Martino LM, Molecular markers for failure of sulfadoxine-pyrimethamine and chlorproguanil-dapsone treatment of Plasmodium falciparum malaria. J Infect Dis. 2002;185:380–8. DOIPubMedGoogle Scholar
- Anderson TJ, Nair S, Jacobzone C, Zavai A, Balkan S. Molecular assessment of drug resistance in Plasmodium falciparum from Bahr El Gazal province, Sudan. Trop Med Int Health. 2003;8:1068–73. DOIPubMedGoogle Scholar
- Nair S, Brockman A, Paiphun L, Nosten F, Anderson TJ. Rapid genotyping of loci involved in antifolate drug resistance in Plasmodium falciparum by primer extension. Int J Parasitol. 2002;32:852–8. DOIPubMedGoogle Scholar
- Nei M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics. 1978;89:583–90.PubMedGoogle Scholar
- Belkhir K, Borsa P, Chikhi L, Raufaste L, Bonhomme F. GENETIX 4.05, Windows software for population genetics (author’s translation) [in French]. Montpellier, France: Laboratoire Génome, Populations, Interactions, CNRS UMR 5000, Université de Montpellier II; 1996–2004 [cited 2010 Sep 5]. http://www.genetix.univ-montp2.fr/genetix/genetix.htm
- Goudet J. FSTAT (version 2.9.3.2), a program to estimate and test population genetics parameters; 2002 [cited 2010 Sep 5]. http://www2.unil.ch/popgen/softwares/fstat.htm
- Wright S. The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution. 1965;19:395–420. DOIGoogle Scholar
- Weir B, Cockerham C. Estimating F-statistics for the analysis of population structure. Evolution. 1984;38:1358–70. DOIGoogle Scholar
- Excoffier L, Laval G, Schneider S. ARLEQUIN (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online. 2005;1:47–50.PubMedGoogle Scholar
- Ter Braack C. Canonical correspondence analysis: a new eigenvector technique for multivariate direct gradient analysis. Ecology. 1986;67:1167–79. DOIGoogle Scholar
- Ter Braack C, Šmilauer P. CANOCO. Fortran program for canonical community ordination [cited 2010 Sep 5]. http://www.pri.wur.nl/UK/products/Canoco/
- Rousset F. Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics. 1997;145:1219–28.PubMedGoogle Scholar
- Durand P, Michalakis Y, Cestier S, Oury B, Leclerc MC, Tibayrenc M, Significant linkage disequilibrium and high genetic diversity in a population of Plasmodium falciparum from an area (Republic of the Congo) highly endemic for malaria. Am J Trop Med Hyg. 2003;68:345–9.PubMedGoogle Scholar
- Razakandrainibe FG, Durand P, Koella JC, De Meeus T, Rousset F, Ayala FJ, “Clonal” population structure of the malaria agent Plasmodium falciparum in high-infection regions. Proc Natl Acad Sci U S A. 2005;102:17388–93. DOIPubMedGoogle Scholar
- Annan Z, Durand P, Ayala FJ, Arnathau C, Awono-Ambene P, Simard F, Population genetic structure of Plasmodium falciparum in the two main African vectors, Anopheles gambiae and Anopheles funestus. Proc Natl Acad Sci U S A. 2007;104:7987–92. DOIPubMedGoogle Scholar
- Bonizzoni M, Afrane Y, Baliraine FN, Amenya DA, Githeko AK, Yan G. Genetic structure of Plasmodium falciparum populations between lowland and highland sites and antimalarial drug resistance in western Kenya. Infect Genet Evol. 2009;9:806–12. DOIPubMedGoogle Scholar
- Ferreira MU, Karunaweera ND, da Silva-Nunes M, da Silva NS, Wirth DF, Hartl DL. Population structure and transmission dynamics of Plasmodium vivax in rural Amazonia. J Infect Dis. 2007;195:1218–26. DOIPubMedGoogle Scholar
- Ariey F, Randrianarivelojosia M, Duchemin JB, Rakotondramarina D, Ouledi A, Robert V, Mapping of a Plasmodium falciparum Pfcrt K76T mutation: a useful strategy for controlling chloroquine resistance in Madagascar. J Infect Dis. 2002;185:710–2. DOIPubMedGoogle Scholar
- Randrianarivelojosia M, Raherinjafy RH, Migliani R, Mercereau-Puijalon O, Ariey F, Bedja SA. Plasmodium falciparum resistant to chloroquine and to pyrimethamine in Comoros. Parasite. 2004;11:419–23.PubMedGoogle Scholar
- Quatresous I, Jeannel D, Sissoko D. Epidémiologie du paludisme à Mayotte. Etat des lieux 2003–2004 et propositions. Paris, France: Institut de Veille Sanitaire; 2005.
- Iyer JK, Milhous WK, Cortese JF, Kublin JG, Plowe CV. Plasmodium falciparum cross-resistance between trimethoprim and pyrimethamine. Lancet. 2001;358:1066–7. DOIPubMedGoogle Scholar
- World Health Organization. Susceptibility of Plasmodium falciparum to antimalarial drugs: report on global monitoring: 1996–2004. Geneva: The Organization; 2005 [cited 2010 Sep 5]. http://www.malaria.org/ABOUT%20MALARIA/SusceptibilityPlasmodium_report%20WHO%202004.pdf
- Talisuna AO, Langi P, Mutabingwa TK, Van Marck E, Speybroeck N, Egwang TG, Intensity of transmission and spread of gene mutations linked to chloroquine and sulphadoxine–pyrimethamine resistance in falciparum malaria. Int J Parasitol. 2003;33:1051–8. DOIPubMedGoogle Scholar
- Hastings IM, Watkins WM. Intensity of malaria transmission and the evolution of drug resistance. Acta Trop. 2005;94:218–29.PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 16, Number 11—November 2010
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Christophe Rogier, UMR 6236–URMITE, IRBA–Antenne de Marseille, Parc du Pharo, BP60109, 13262 Marseille CEDEX 07, France
Top