Volume 23, Number 2—February 2017
CME ACTIVITY - Research
Multidrug-Resistant Candida haemulonii and C. auris, Tel Aviv, Israel
Cite This Article
Citation for Media
Introduction

This activity has been planned and implemented through the joint providership of Medscape, LLC and Emerging Infectious Diseases. Medscape, LLC is accredited by the American Nurses Credentialing Center (ANCC), the Accreditation Council for Pharmacy Education (ACPE), and the Accreditation Council for Continuing Medical Education (ACCME), to provide continuing education for the healthcare team.
Medscape, LLC designates this Journal-based CME activity for a maximum of 1.00 AMA PRA Category 1 Credit(s)™. Physicians should claim only the credit commensurate with the extent of their participation in the activity.
All other clinicians completing this activity will be issued a certificate of participation. To participate in this journal CME activity: (1) review the learning objectives and author disclosures; (2) study the education content; (3) take the post-test with a 75% minimum passing score and complete the evaluation at http://www.medscape.org/journal/eid; and (4) view/print certificate.
Release date: January 13, 2017; Expiration date: January 13, 2018
Learning Objectives
Upon completion of this activity, participants will be able to:
1. Assess the phylogenetic analysis of Candida auris and C. haemulonii in the current study
2. Analyze patient characteristics among infections with C. auris and C. haemulonii
3. Distinguish the pattern of antifungal resistance in infections with C. auris and C. haemulonii
4. Evaluate the pathologic potential of C. auris and C. haemulonii
CME Editor
Karen L. Foster, MA, Technical Writer/Editor, Emerging Infectious Diseases. Disclosure: Karen L. Foster has disclosed no relevant financial relationships.
CME Author
Charles P. Vega, MD, Clinical Professor of Family Medicine, University of California, Irvine. Disclosure: Charles P. Vega, MD, has disclosed the following financial relationships: served as an advisor or consultant for Allergan, Inc.; McNeil Consumer Healthcare; served as a speaker or a member of a speakers bureau for Shire Pharmaceuticals.
Authors
Disclosures: Ronen Ben-Ami, MD, has disclosed the following relevant financial relationships: served as an advisor or consultant for Pfizer, Inc.; Merck & Co. Inc.; Neopharm; Teva Pharmaceuticals. Yasmin Maor, MD, has disclosed the following relevant financial relationships: served as an advisor or consultant for Novo Nordisk; served as a speaker or member of a speakers bureau for Pfizer; MSD. Judith Berman, PhD; Ana Novikov, MSc; Edna Bash, MSc; Yael Shachor-Meyouhas, MD; Shiri Zakin, PhD; Jalal Tarabia, PhD; Vered Schechner, MD, MSc; Amos Adler, MD; and Talya Finn, MBBS, have disclosed no relevant financial relationships.
Abstract
Candida auris and C. haemulonii are closely related, multidrug-resistant emerging fungal pathogens that are not readily distinguishable with phenotypic assays. We studied C. auris and C. haemulonii clinical isolates from 2 hospitals in central Israel. C. auris was isolated in 5 patients with nosocomial bloodstream infection, and C. haemulonii was found as a colonizer of leg wounds at a peripheral vascular disease clinic. Liberal use of topical miconazole and close contact among patients were implicated in C. haemulonii transmission. C. auris exhibited higher thermotolerance, virulence in a mouse infection model, and ATP-dependent drug efflux activity than C. haemulonii. Comparison of ribosomal DNA sequences found that C. auris strains from Israel were phylogenetically distinct from isolates from East Asia, South Africa and Kuwait, whereas C. haemulonii strains from different countries were closely interrelated. Our findings highlight the pathogenicity of C. auris and underscore the need to limit its spread.
Candida species are leading causes of bloodstream infection (BSI) in hospitalized patients, particularly those in intensive care units who are exposed to broad-spectrum antimicrobial drugs, indwelling vascular catheters, parenteral nutrition, abdominal surgery, and immunosuppressive agents (1,2). High rates of attributable death have been associated with delayed initiation of appropriate antifungal treatment (3,4). This problem is compounded by the emergence of drug-resistant Candida species, notably C. glabrata, in many hospitals (5).
C. auris is an emerging opportunistic pathogen, first reported in 2009 as an isolate from the external ear of an inpatient at a hospital in Japan (6). It has since been identified as a cause of nosocomial BSI in numerous countries in East Asia, the Middle East, Africa, and Europe (7–11). C. auris might be resistant to multiple classes of antifungal agents and apparently has a potential for person-to-person transmission, challenging clinicians and infection control teams (12). C. auris often is misidentified by traditional microbiological methods as C. haemulonii, a phylogenetically related drug-resistant Candida species that also is increasingly reported in healthcare facilities worldwide (13).
We report on the detection of multidrug-resistant C. auris and C. haemulonii in clinical specimens in Tel Aviv, Israel, and specifically on the emergence of C. auris as a cause of nosocomial BSI. We highlight distinct clinical and epidemiologic characteristics of these 2 species and present experimental evidence for differences in their virulence.
We undertook this study after C. auris BSI was detected in 4 patients during May–October 2014 at the Tel Aviv Sourasky Medical Center (TASMC), a tertiary-level hospital in Tel Aviv. An additional C. auris bloodstream isolate was recovered in April 2015 from a patient at the Wolfson Medical Center in Holon (southern Tel Aviv metropolitan area). No additional C. haemulonii or C. auris isolates were identified through inquiries at additional clinical microbiology laboratories in Israel.
The TASMC Institutional ethics committee approved this study. Need for informed consent was waived because of the observational and anonymous nature of the study.
Clinical Candida isolates
Candida isolates recovered from clinical specimens were identified at the TASMC Clinical Microbiology Laboratory by growth characteristics on CHROMagar Candida (CHROMagar, Paris, France) and the Vitek 2 YST ID system (bioMérieux, Marcy-l’Étoile, France). The Vitek 2 database does not include C. auris, and this species is routinely misidentified as C. haemulonii (13). We therefore reviewed all isolates identified as C. haemulonii during January 2009–August 2015. Isolates recovered during May 2014–August 2015 were stored at −20°C and subjected to further analyses. We assessed thermotolerance by plating serial dilutions of yeast culture on Sabouraud dextrose agar (SDA) plates and assessing growth after 24 h incubation at 35°C–42°C.
Sequence-Based Species Identification
Candida isolates were streaked on SDA plates to ensure purity. We extracted DNA by using PrepMan Ultra solution (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s instructions and amplified and sequenced the internal transcribed spacer (ITS) and D1/D2 large subunit (LSU) ribosomal DNA segments by using primer pairs ITS1/ITS4 and LSU1/LSU2 (14), respectively. PCR was performed in 0.2-mL tubes with 0.4 μmol/L or 0.2 μmol/L of each primer for ITS and LSU, respectively; 10 μL Larova Red Load Taq Master Mix (5×) (Larova, Jena, Germany); and ≈25 ng of template. PCR conditions were 95°C for 4.5 min (denaturation), 40 cycles of 95°C for 30 s (denaturation), 55°C (ITS) or 48°C (LSU) for 30 s (annealing), 72°C for 1 min (extension), and a final extension stage of 72°C for 7 min. PCR products were resolved on 0.7% agarose gel and stained with SERVA DNA stain clear G (Tamar, Mevaseret Zion, Israel). Products were cleaned with QIAquick PCR purification kit (QIAGEN, Hilden, Germany) and sequenced at Hy-Labs (Rehovot, Israel). We then aligned ITS and LSU sequences with the matching type strain sequences for CBS5149T (C. haemulonii), CBS7798T (C. duobushaemulonii), CNM-CL7239T (C. haemulonii var. vulnera), CBS10099T (C. pseudohaemulonii), and CBS10913T (C. auris). A similarity score of >98% in both ITS and LSU sequences was required for species-level identification. All new sequences were deposited in GenBank (Table 1).
Phylogenetic Analyses
We aligned ITS and LSU sequences of C. haemulonii and C. auris isolates by using MUSCLE (15) and generated phylogenetic trees with the neighbor-joining method (16), using the Kimura 2-parameter method to compute evolutionary distances (17). We tested phylogeny with the bootstrap method (500 replicates) and used Schizosaccharomyces pombe strains ATCC 38366 and CBS 356 as outgroups. Evolutionary analyses were performed in MEGA7 (18).
Patient Characteristics
We retrospectively reviewed the medical records of patients from whom Vitek-identified C. haemulonii was recovered from any site and recorded patient demographics, hospital unit, co-morbidities, medications, and clinical characteristics by using a structured form. We found 40 patient-specific C. haemulonii isolates, 20 (50%) of which originated from patients receiving care at an outpatient peripheral vascular disease clinic (clinic A). We therefore conducted an investigation at clinic A, which included review of patient treatment protocols, observed patient care, and surveillance mycologic cultures from environmental surfaces, wound irrigation solutions, dressings, and the hands of medical staff. To define risk factors for C. haemulonii colonization, we conducted an unmatched case–control study using 40 noncolonized patients followed at clinic A as controls.
Antifungal Susceptibility Testing
We determined MICs of fluconazole, itraconazole, voriconazole, posaconazole, amphotericin B, anidulafungin, micafungin, caspofungin, and flucytosine by broth microdilution using Clinical and Laboratory Standards Institute methods (19). Results were read after 48 h for azoles, amphotericin B, and flucytosine and after 24 h for echinocandins.
Rhodamine 6G Efflux
To assess ABC-type drug transporter activity, we determined glucose-induced efflux of rhodamine 6G, as described previously (20,21). We grew Candida isolates to log-phase in liquid yeast extract glucose at 35°C. We then collected yeast cells by centrifugation, transferred 109 cells to 20 mL fresh yeast extract glucose, and incubated them at 27°C for an additional 2 h. Next, we collected yeast cells by centrifugation, washed them twice in phosphate-buffered saline (PBS), and added 10 mL PBS containing 15 μM rhodamine 6G without glucose to the pellets. Suspensions were vortexed and incubated at 27°C for 90 min to enable rhodamine 6G uptake under carbon source–depleted conditions. We then collected cells by centrifugation, washed them twice in PBS, and suspended them in 750 μL PBS in microfuge tubes. To start rhodamine 6G efflux, we added 250 μL PBS with 8 mmol/L glucose. We prepared control tubes with glucose-free PBS, removed them after 5, 15, and 25 min of incubation at 35°C, and measured fluorescence in 200-μL aliquots of supernatant by using a spectrophotometer at excitation 527 nm and emission 555 nm (Synergy HT, BioTek, Winooski, VT, USA).
Mouse Model of Disseminated Candidiasis
To assess the relative virulence of C. haemulonii and C. auris, we determined the lethality and tissue fungal loads of representative strains in a mouse model of hematogenous disseminated candidiasis. Experiments were approved by the TASMC Institutional Animal Care and Use Ethics Committee. We used cyclophosphamide (150 mg/kg intraperitoneally) to immunosuppress 6-week-old female BALB/c mice weighing 16–20 g (Harlan, Rehovot, Israel) 3 days before and on the day of infection. Candida cells were collected from log-phase culture on the day of infection and washed twice in sterile PBS. Mice were infected in groups of 10 with C. haemulonii strain TA001-14, C. auris strain TA005-14, and C. albicans strain CBS 8837. We injected 100 μL of PBS containing 7 × 107 yeast cells intravenously into the lateral tail vein of each animal. A control group received intravenous injection of cell-free PBS. Death was assessed over 12 days. Kidney tissue fungal loads were determined in separate experiments where mice were similarly immunosuppressed and infected intravenously with 4 × 107 yeast cells/100 μL PBS. Seven days after infection, mice were killed by CO2 inhalation, and kidneys were excised aseptically, weighed, and homogenized in a TissueLyser (QIAGEN). Homogenates were serially diluted 10- to 1,000-fold in sterile saline and plated on SDA. We calculated fungal loads (CFU per gram of tissue) from colony counts after 48 h incubation at 35°C.
Statistical Analyses
We compared continuous variables between case and control patients using the Student t test for normally distributed variables and the Wilcoxon rank-sum test for non–normally distributed variables. We compared dichotomous variables using Fisher’s exact test. Rhodamine 6G efflux, expressed as relative fluorescence units, was computed for each Candida strain and compared by using 1-way analysis of variance. We used Dunnett’s multiple comparisons test to compare relative fluorescence unit values of specific C. haemulonii and C. auris strains with averaged control values of 5 C. glabrata strains. Survival curves of mice infected with different Candida strains were plotted by using the Kaplan-Meier method and compared with the log-rank test. We considered 2-tailed p values <0.05 statistically significant.
Sequence-Based Identification
We identified 40 patient-specific Candida strains as C. haemulonii by the Vitek-2 YST ID system during January 2009–July 2015. Isolates were recovered from wounds (n = 24), urine (n = 9), blood (n = 5), and central venous catheter tips (n = 2). Of these, 9 isolates recovered during May 2014–May 2015 were available for analysis; we identified 6 (including the 5 blood isolates) as C. auris and 3 as C. haemulonii by ITS and LSU sequencing (Table 1). Sequences were 100% identical among strains of each species. C. auris strains from Israel shared 98.6% and 98.3% similarity of ITS and LSU sequences, respectively, with the C. auris type strain CBS10913T. C. haemulonii strains were 100% identical to C. haemulonii CBS5149T on the basis of ITS and LSU sequences.
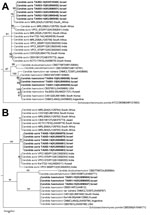
Phylogenetic trees based on ITS and LSU sequences showed that the C. auris isolates from Tel Aviv are distinct from other isolates from East Asia, Africa, and the Middle East. Specifically, isolates from Israel showed 98.6% similarity of ITS and LSU sequences with the India clone, represented by CBS12768, 96.2% similarity with the South Korea clone, and 96.7% similarity with strain CH1 from Kuwait. In contrast, ITS and LSU sequences from Israel C. haemulonii strains were 100% homologous with C. haemulonii from South Korea, Brazil, and Kuwait, suggesting worldwide predominance of a single C. haemulonii clone (Figure 1).
Clinical Features
Eight of 9 patients with sequence-validated isolates were hospitalized at TASMC (Table 1). An additional patient with C. auris infection was hospitalized at the Wolfson Medical Center, but was receiving regular care for HIV infection at TASMC. All 3 C. haemulonii isolates were recovered from chronic leg ulcers of patients with peripheral vascular disease, 2 of whom were treated at vascular outpatient clinic A. Five of 6 C. auris isolates represented BSI: 3 patients had vascular catheter–related candidemia, and 2 had primary nosocomial candidemia of unclear origin. Two of 5 patients with C. auris BSI died during hospitalization.
We reviewed the medical records of 40 patients with Vitek-identified C. haemulonii cultures. Thirty-three (83%) were male. Median age was 74 years (range 37–91 years). Nineteen (48%) had peripheral vascular disease, 20 (50%) had diabetes mellitus, 22 (55%) had ischemic heart disease, and 11 (28%) had end-stage renal disease. Twenty patients (50%) were receiving regular care at clinic A, representing 8% (20/261) of all clinic patients. In all 20 patients, C. haemulonii had been recovered from chronic leg ulcers, and none had documented wound infection at the time of culture. Cultures of environmental surfaces, medical devices, dressings, irrigation solutions, and hands of medical staff were negative for yeast. Compared with 40 control patients who were not carriers of C. haemulonii, carriers were older, had a lower glomerular filtration rate, and were more likely to be male and to have ischemic heart disease (Table 2). Observations revealed a practice among medical staff of routinely applying topical miconazole cream to chronic ulcers without evidence of infection. Periodic wound cultures were obtained regularly, irrespective of signs of ulcer inflammation or purulence. Multiple social interactions were noted among patients in a single room where wound care was performed.
Antifungal Susceptibility
All C. haemulonii and C. auris isolates had fluconazole MICs >8 mg/L (range 16–64 mg/L; MIC50 32 mg/L). MICs of other azoles were also elevated: itraconazole, 0.25 to >16 mg/L (MIC50 0.5 mg/L); voriconazole, 0.25–1 mg/L (MIC50 0.5 mg/L); and posaconazole, 0.06 to >8 mg/L (MIC50 0.25 mg/L). Amphotericin B MIC ranged from 1 to 2 mg/L for C. auris isolates and from 2 to 8 mg/L for C. haemulonii isolates. All isolates appeared susceptible to anidulafungin (MIC 0.03 mg/L) and all isolates except 1 C. haemulonii were susceptible to micafungin (MIC 0.12–0.5 mg/L; MIC50 0.12 mg/L). Caspofungin MIC was 0.5 mg/L for all isolates. All isolates except 1 C. auris were susceptible to flucytosine (Table 3).
Rhodamine 6G Efflux
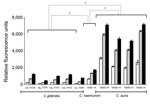
Rhodamine 6G is a substrate of ATP binding cassette (ABC) type efflux pumps responsible for multiazole resistance in C. glabrata. C. haemulonii and C. auris strains exhibited robust rhodamine 6G efflux activity when glucose (8 mM) was present in the medium, consistent with ABC-type transport. Rhodamine 6G efflux of C. auris strains was significantly greater than that of C. glabrata strains (14.4-, 10-, and 6.7-fold higher at 5, 15, and 25 min, respectively; p<0.0001) and C. haemulonii (3.8-, 3.8-, and 3.6-fold higher at 5, 15, and 25 min, respectively; p<0.0001). C. haemulonii showed greater rhodamine 6G efflux than C. glabrata (3.8-, 2.7-, and 1.9-fold higher at 5, 15, and 25 min, respectively (p<0.0001) (Figure 2).
Thermotolerance
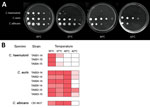
Survival and growth at physiologic temperature are prerequisites for microbial invasion and pathogenicity. C. haemulonii isolates grew well at 35°C, but growth at 37°C was poor or absent, and no growth occurred at 40°C and 42°C. In contrast, growth of C. auris isolates at 37°C and 40°C was similar to that of C. albicans, and 4 of 6 isolates grew at 42°C (Figure 3).
Virulence in a Mouse Model of Disseminated Candidiasis
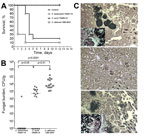
We compared the virulence of C. auris and C. haemulonii isolates in a mouse model of hematogenous disseminated candidiasis. C. haemulonii was completely nonvirulent in this model; 100% of mice survived 12 days after inoculation with no visible signs of illness. In contrast, inoculation with C. auris resulted in rapid death and only 20% survival 5 days after infection (p = 0.0002, log-rank test). Death of mice infected with C. auris was significantly less rapid than that of mice infected with C. albicans (median survival 4 d and 1 d, respectively; p = 0.01; Figure 4, panel A). Kidney tissue fungal load correlated with survival rates. Specifically, we recovered no viable yeast cells from kidneys of mice inoculated with C. haemulonii, whereas infection with C. auris and C. albicans yielded median tissue loads of 5.9 × 104 CFU/g and 7.1 × 105 CFU/g, respectively (p<0.0001; Figure 4, panel B). Histopathologic analysis showed yeast cell aggregates in kidneys of C. auris–inoculated mice, distinct from tissue invasive hyphae observed in C. albicans–infected kidneys (Figure 4, panel C).
Concern about the international emergence and spread of C. auris as a cause of invasive infection in hospitals stems from 3 characteristics of this opportunistic pathogen (12,13): 1) resistance to multiple antifungal drugs and possibly to all major classes of systemic antifungal drugs; 2) horizontal transmission among hospitalized patients, leading to nosocomial outbreaks (8,10,11,13); and 3) high associated death rates (7,8,10). C. auris and C. haemulonii are phylogenetically related species in the Metschnikowia clade that share a propensity for multidrug resistance. We identified C. auris and C. haemulonii in 2 hospitals in Israel and highlighted clinical and experimental evidence for differences in the drug-susceptibility patterns, drug efflux activity, pathogenicity, and global phylogenetics of these 2 species.
In our study, C. auris and C. haemulonii had high MICs of azoles and amphotericin B. Echinocandin MICs were within the susceptible range. An amphotericin B epidemiologic cutoff value of 2 mg/L previously was established (23), but clinical correlation between amphotericin B MIC and treatment outcomes is lacking (24). Compared with C. auris, C. haemulonii isolates had higher amphotericin B MICs. The relevance of these resistance patterns to treatment strategies remains to be determined.
ABC-type efflux activity, as evidenced by Rhodamine 6G transport, was significantly greater among C. auris than C. glabrata isolates. This observation provides a mechanistic basis for the intrinsic resistance of C. auris to azoles and is consistent with the identification of multiple putative transporter-encoding genes belonging to the ABC and major facilitator gene families in the C. auris genome (25).
Of Vitek-identified C. haemulonii isolates at TASMC, 50% were wound cultures from patients cared for at clinic A. That most of these isolates were not available for sequencing is a limitation of our study. However, we identified the 2 Candida isolates from clinic A patients that were available by ITS and LSU sequencing as C. haemulonii, and all 3 sequence-identified C. haemulonii isolates were recovered from leg ulcers of patients with peripheral vascular disease. Colonization of patients treated in close proximity in 1 room strongly suggests person-to-person transmission and supports interim guidelines for contact isolation (26). However, we were unable to identify an environmental reservoir of C. haemulonii. We suggest that topical application of miconazole to wounds most likely caused selective pressure and facilitated the overgrowth of C. haemulonii. After this investigation and termination of routine topical azole use, no additional cases of C. haemulonii were detected in clinic A during April 2015–July 2016.
We recovered 5 of 6 sequence-identified C. auris isolates from patients with nosocomial BSI. In contrast, all C. haemulonii isolates were cultured from superficial wounds. This observation reflects the global epidemiology of these species. C. haemulonii has been isolated from chronic leg ulcers of patients in India and Brazil (27,28). C. auris has caused outbreaks of BSI in the United Kingdom (13), India (8), Kenya (11), South Africa (9), and South Korea (29), whereas reports of C. haemulonii as an agent of BSI have been infrequent (27,29–32). Moreover, C. auris fungemia is associated with high death rates (8,10), contrasting with reports of patients surviving prolonged C. haemulonii fungemia (31). Fatal C. haemulonii fungemia, although rare, has been reported in neonates and in patients with cancer and neutropenia (27,32).
In our study, C. auris, but not C. haemulonii, grew at 37°C–42°C and exhibited lethality and tissue invasion in a mouse model of invasive candidiasis only slightly less than those of C. albicans, the prototypical pathogenic Candida species. Both C. auris and C. haemulonii are unable to form hyphae, which contribute to virulence in C. albicans. Formation of large aggregates resulting from failure of budding yeast to separate has been noted in some C. auris isolates (33). We observed distinct yeast cell aggregates in the kidneys of mice with lethal C. auris infection, which suggests that aggregation might be a mode of immune evasion and persistence in tissue. The C. auris genome contains C. albicans gene orthologs, such as secreted proteinases and mannosyl transferases, which might have roles in pathogenesis (25). However, C. auris has a genome that is highly divergent from those of other Candida species, and most of its genes have not yet been characterized (25).
Ribosomal DNA sequences were identical among C. haemulonii strains from Israel, Kuwait, East Asia, South America, and the United States. In contrast, the global phylogenetics of C. auris demonstrate distinct clones for each country, indicating greater genomic diversity for this species. Further study is needed to establish whether the divergence of C. auris clones translates into country-specific patterns of invasiveness, virulence, and drug resistance. Our findings affirm the need for intensified vigilance and mobilization of infection control measures to limit the spread of C. auris.
Dr. Ben-Ami is head of the Infectious Diseases Unit and the National Center for Clinical Mycology at the Tel Aviv Medical Center, Israel. His research focuses on the epidemiology, drug resistance traits, and virulence determinants of medically important fungi.
Acknowledgment
J.B. was supported by the European Research Council Advanced Award 340087 (RAPLODAPT).
References
- Magill SS, Edwards JR, Bamberg W, Beldavs ZG, Dumyati G, Kainer MA, et al.; Emerging Infections Program Healthcare-Associated Infections and Antimicrobial Use Prevalence Survey Team. Multistate point-prevalence survey of health care-associated infections. N Engl J Med. 2014;370:1198–208. DOIPubMedGoogle Scholar
- Kullberg BJ, Arendrup MC. Invasive Candidiasis. N Engl J Med. 2015;373:1445–56. DOIPubMedGoogle Scholar
- Garey KW, Rege M, Pai MP, Mingo DE, Suda KJ, Turpin RS, et al. Time to initiation of fluconazole therapy impacts mortality in patients with candidemia: a multi-institutional study. Clin Infect Dis. 2006;43:25–31. DOIPubMedGoogle Scholar
- Andes DR, Safdar N, Baddley JW, Playford G, Reboli AC, Rex JH, et al.; Mycoses Study Group. Impact of treatment strategy on outcomes in patients with candidemia and other forms of invasive candidiasis: a patient-level quantitative review of randomized trials. Clin Infect Dis. 2012;54:1110–22. DOIPubMedGoogle Scholar
- Alexander BD, Johnson MD, Pfeiffer CD, Jiménez-Ortigosa C, Catania J, Booker R, et al. Increasing echinocandin resistance in Candida glabrata: clinical failure correlates with presence of FKS mutations and elevated minimum inhibitory concentrations. Clin Infect Dis. 2013;56:1724–32. DOIPubMedGoogle Scholar
- Satoh K, Makimura K, Hasumi Y, Nishiyama Y, Uchida K, Yamaguchi H. Candida auris sp. nov., a novel ascomycetous yeast isolated from the external ear canal of an inpatient in a Japanese hospital. Microbiol Immunol. 2009;53:41–4. DOIPubMedGoogle Scholar
- Lee WG, Shin JH, Uh Y, Kang MG, Kim SH, Park KH, et al. First three reported cases of nosocomial fungemia caused by Candida auris. J Clin Microbiol. 2011;49:3139–42. DOIPubMedGoogle Scholar
- Chowdhary A, Sharma C, Duggal S, Agarwal K, Prakash A, Singh PK, et al. New clonal strain of Candida auris, Delhi, India. Emerg Infect Dis. 2013;19:1670–3. DOIPubMedGoogle Scholar
- Magobo RE, Corcoran C, Seetharam S, Govender NP. Candida auris-associated candidemia, South Africa. Emerg Infect Dis. 2014;20:1250–1. DOIPubMedGoogle Scholar
- Chowdhary A, Anil Kumar V, Sharma C, Prakash A, Agarwal K, Babu R, et al. Multidrug-resistant endemic clonal strain of Candida auris in India. Eur J Clin Microbiol Infect Dis. 2014;33:919–26. DOIPubMedGoogle Scholar
- Okinda N, Kagotho E, Castanheira M, Njuguna A, Omuse G, Makau P, et al. Candidemia at a referral hospital in sub-Saharan Africa: emergence of Candida auris as a major pathogen. European Conference on Clinical Microbiology and Infectious Diseases; 2014 May 10–13; Barcelona, Spain.
- Centers for Disease Control and Prevention. Global emergence of invasive infections caused by the multidrug-resistant yeast Candida auris [cited 2016 Jul 29]. http://www.cdc.gov/fungal/diseases/candidiasis/candida-auris-alert.html
- Public Health England. Candida auris identified in England [cited 2016 Jul 1]. https://www.gov.uk/government/publications/candida-auris-emergence-in-england/candida-auris-identified-in-england
- Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, et al.; Fungal Barcoding Consortium; Fungal Barcoding Consortium Author List. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A. 2012;109:6241–6. DOIPubMedGoogle Scholar
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7. DOIPubMedGoogle Scholar
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.PubMedGoogle Scholar
- Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111–20. DOIPubMedGoogle Scholar
- Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4. DOIPubMedGoogle Scholar
- Clinical and Laboratory Standards Institute. Reference method for broth dilution antifungal susceptibility testing of yeasts. Approved standard M27-A3. Wayne (PA): The Institute; 2008.
- Maesaki S, Marichal P, Vanden Bossche H, Sanglard D, Kohno S. Rhodamine 6G efflux for the detection of CDR1-overexpressing azole-resistant Candida albicans strains. J Antimicrob Chemother. 1999;44:27–31. DOIPubMedGoogle Scholar
- Kolaczkowski M, van der Rest M, Cybularz-Kolaczkowska A, Soumillion JP, Konings WN, Goffeau A. Anticancer drugs, ionophoric peptides, and steroids as substrates of the yeast multidrug transporter Pdr5p. J Biol Chem. 1996;271:31543–8. DOIPubMedGoogle Scholar
- Levey AS, Coresh J, Greene T, Stevens LA, Zhang YL, Hendriksen S, et al.; Chronic Kidney Disease Epidemiology Collaboration. Using standardized serum creatinine values in the modification of diet in renal disease study equation for estimating glomerular filtration rate. Ann Intern Med. 2006;145:247–54. DOIPubMedGoogle Scholar
- Pfaller MA, Espinel-Ingroff A, Canton E, Castanheira M, Cuenca-Estrella M, Diekema DJ, et al. Wild-type MIC distributions and epidemiological cutoff values for amphotericin B, flucytosine, and itraconazole and Candida spp. as determined by CLSI broth microdilution. J Clin Microbiol. 2012;50:2040–6. DOIPubMedGoogle Scholar
- Park BJ, Arthington-Skaggs BA, Hajjeh RA, Iqbal N, Ciblak MA, Lee-Yang W, et al. Evaluation of amphotericin B interpretive breakpoints for Candida bloodstream isolates by correlation with therapeutic outcome. Antimicrob Agents Chemother. 2006;50:1287–92. DOIPubMedGoogle Scholar
- Chatterjee S, Alampalli SV, Nageshan RK, Chettiar ST, Joshi S, Tatu US. Draft genome of a commonly misdiagnosed multidrug resistant pathogen Candida auris. BMC Genomics. 2015;16:686. DOIPubMedGoogle Scholar
- Public Health England. Guidance for the laboratory investigation, management and infection prevention and control for cases of Candida auris [cited 2016 Jul 1]. https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/534174/Guidance_Candida__auris.pdf
- de Almeida JN Jr, Assy JG, Levin AS, Del Negro GM, Giudice MC, Tringoni MP, et al. Candida haemulonii complex species, Brazil, January 2010–March 2015. Emerg Infect Dis. 2016;22:561–3. DOIPubMedGoogle Scholar
- Kumar A, Prakash A, Singh A, Kumar H, Hagen F, Meis JF, et al. Candida haemulonii species complex: an emerging species in India and its genetic diversity assessed with multilocus sequence and amplified fragment-length polymorphism analyses. Emerg Microbes Infect. 2016;5:e49. DOIPubMedGoogle Scholar
- Kim MN, Shin JH, Sung H, Lee K, Kim EC, Ryoo N, et al. Candida haemulonii and closely related species at 5 university hospitals in Korea: identification, antifungal susceptibility, and clinical features. Clin Infect Dis. 2009;48:e57–61. DOIPubMedGoogle Scholar
- Ramos LS, Figueiredo-Carvalho MH, Barbedo LS, Ziccardi M, Chaves AL, Zancopé-Oliveira RM, et al. Candida haemulonii complex: species identification and antifungal susceptibility profiles of clinical isolates from Brazil. J Antimicrob Chemother. 2015;70:111–5. DOIPubMedGoogle Scholar
- Ruan SY, Kuo YW, Huang CT, Hsiue HC, Hsueh PR. Infections due to Candida haemulonii: species identification, antifungal susceptibility and outcomes. Int J Antimicrob Agents. 2010;35:85–8. DOIPubMedGoogle Scholar
- Khan ZU, Al-Sweih NA, Ahmad S, Al-Kazemi N, Khan S, Joseph L, et al. Outbreak of fungemia among neonates caused by Candida haemulonii resistant to amphotericin B, itraconazole, and fluconazole. J Clin Microbiol. 2007;45:2025–7. DOIPubMedGoogle Scholar
- Borman AM, Szekely A, Johnson EM, Mitchell AP. Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic Candida species. mSphere. 2016;1.pii:e00189-16. PubMedGoogle Scholar
Figures
Tables
Follow Up
Earning CME Credit
To obtain credit, you should first read the journal article. After reading the article, you should be able to answer the following, related, multiple-choice questions. To complete the questions (with a minimum 75% passing score) and earn continuing medical education (CME) credit, please go to http://www.medscape.org/journal/eid. Credit cannot be obtained for tests completed on paper, although you may use the worksheet below to keep a record of your answers. You must be a registered user on Medscape.org. If you are not registered on Medscape.org, please click on the “Register” link on the right hand side of the website to register. Only one answer is correct for each question. Once you successfully answer all post-test questions you will be able to view and/or print your certificate. For questions regarding the content of this activity, contact the accredited provider, CME@medscape.net. For technical assistance, contact CME@webmd.net. American Medical Association’s Physician’s Recognition Award (AMA PRA) credits are accepted in the US as evidence of participation in CME activities. For further information on this award, please refer to http://www.ama-assn.org/ama/pub/about-ama/awards/ama-physicians-recognition-award.page. The AMA has determined that physicians not licensed in the US who participate in this CME activity are eligible for AMA PRA Category 1 Credits™. Through agreements that the AMA has made with agencies in some countries, AMA PRA credit may be acceptable as evidence of participation in CME activities. If you are not licensed in the US, please complete the questions online, print the certificate and present it to your national medical association for review.
Article Title: Multidrug-Resistant Candida haemulonii and C. auris, Tel Aviv, Israel
CME Questions
1. Which of the following statements regarding the phylogenetic trees of Candida auris and C. haemulonii in the current study is most accurate?
A. Both Candida strains were closely linked with singular global international clones
B. C. auris was distinct from other global isolates; C. haemulonii was similar to a single global isolate
C. C. auris was similar to a single global isolate; C. haemulonii was distinct from other global isolates
D. Both Candida strains were distinct from other global isolates
2. Which of the following statements regarding patient characteristics in the current study of C. auris and C. haemulonii infection is most accurate?
A. Most cases of C. auris infection were isolated from skin samples
B. Carriers of C. haemulonii tended to be younger and healthier compared with noncarriers
C. C. haemulonii was most frequently recovered from sites of healed lower extremity ulcers
D. Cultures of medical personnel and equipment implicated chronic facility colonization with C. haemulonii
3. C. auris and C. haemulonii were most likely to be resistant to which of the following antifungals in the current study?
A. Fluconazole
B. Anidulafungin
C. Micafungin
D. Flucytosine
4. What else did the current study demonstrate regarding the pathologic potential of C. auris and C. haemulonii?
A. C. auris demonstrated greater potential for azole resistance compared with C. haemulonii
B. The potential for azole resistance in C. auris and C. haemulonii was not as strong as that of C. glabrata
C. Only C. auris grew poorly at high temperatures
D. Both C. auris and C. haemulonii grew poorly at high temperatures
Activity Evaluation
|
1. The activity supported the learning objectives. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
2. The material was organized clearly for learning to occur. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
3. The content learned from this activity will impact my practice. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
4. The activity was presented objectively and free of commercial bias. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
Related Links
Table of Contents – Volume 23, Number 2—February 2017
| EID Search Options |
|---|
|
|
|
|
|
|



Please use the form below to submit correspondence to the authors or contact them at the following address:
Ronen Ben-Ami, Infectious Diseases Unit, Tel Aviv Sourasky Medical Center, 6 Weizmann St, Tel Aviv 64239, Israel
Top