Volume 32, Number 3—March 2026
Dispatch
Extraintestinal Entamoeba moshkovskii Infection, Eastern India
Figure 3
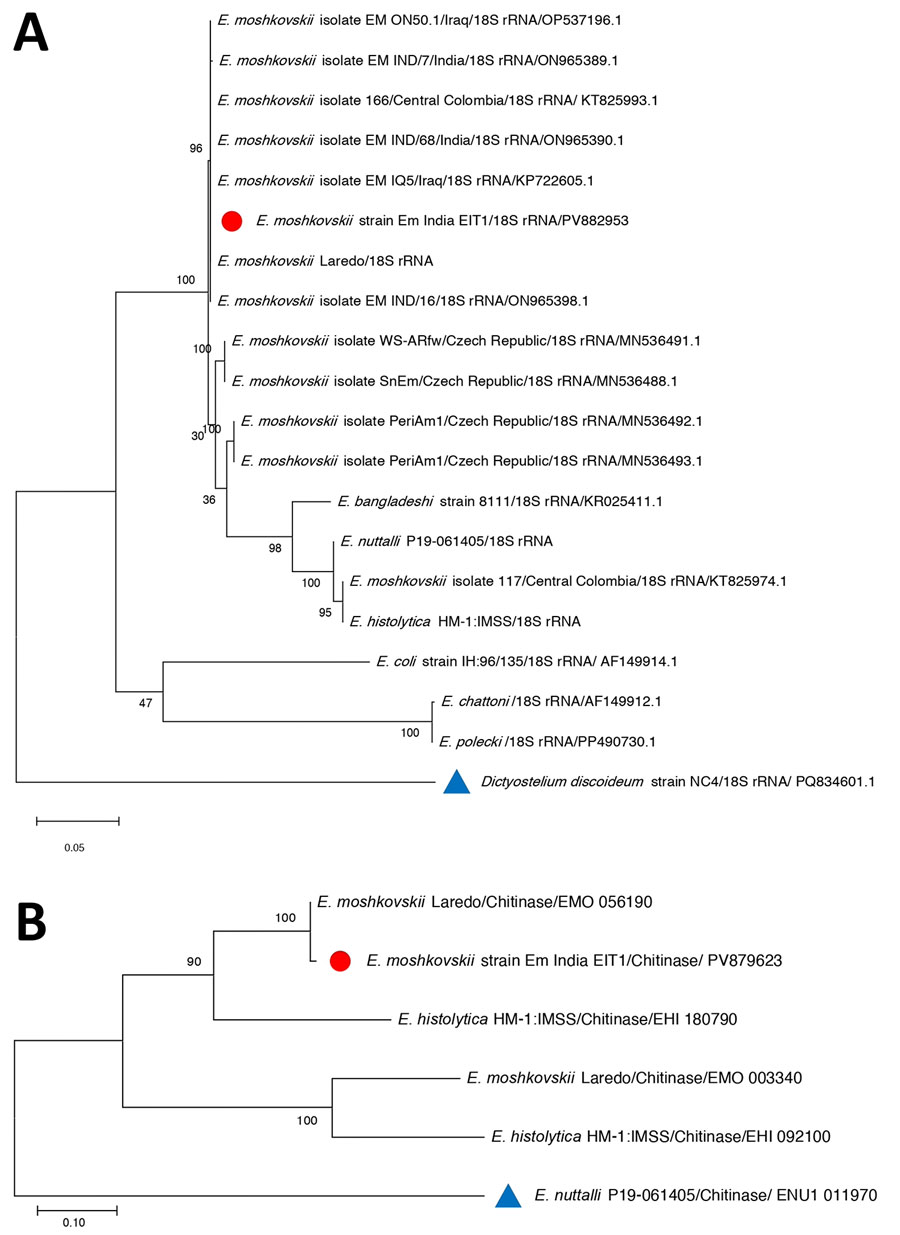
Figure 3. Phylogenetic trees constructed from the 18S rRNA and chitinase gene sequences obtained from Entamoeba moshkovskii isolates from a patient in eastern India (red circles) and reference sequences. A) Phylogenetic tree of 18S rRNA. The tree with the highest log-likelihood value (–2,562.32) is shown. Initial heuristic trees were generated automatically using the maximum parsimony approach. The model incorporated a proportion of invariant sites (45.63%). The analysis included 9 nucleotide sequences with 859 aligned positions in the final dataset. B) Phylogenetic tree of chitinase gene sequence. The tree with the highest log-likelihood value (−3,301.96) is presented. Initial heuristic searches were performed automatically using the maximum parsimony method. The final analysis included 20 nucleotide sequences, with a total of 939 positions in the final dataset. GenBank accession numbers are provided. Outgroup taxa are indicated with blue triangles. The phylogeny was inferred using the maximum likelihood method. Scale bars indicate substitutions per site.
1These authors contributed equally to this article.