Volume 23, Number 2—February 2017
Research
Swine Influenza Virus (H1N2) Characterization and Transmission in Ferrets, Chile
Figure 2
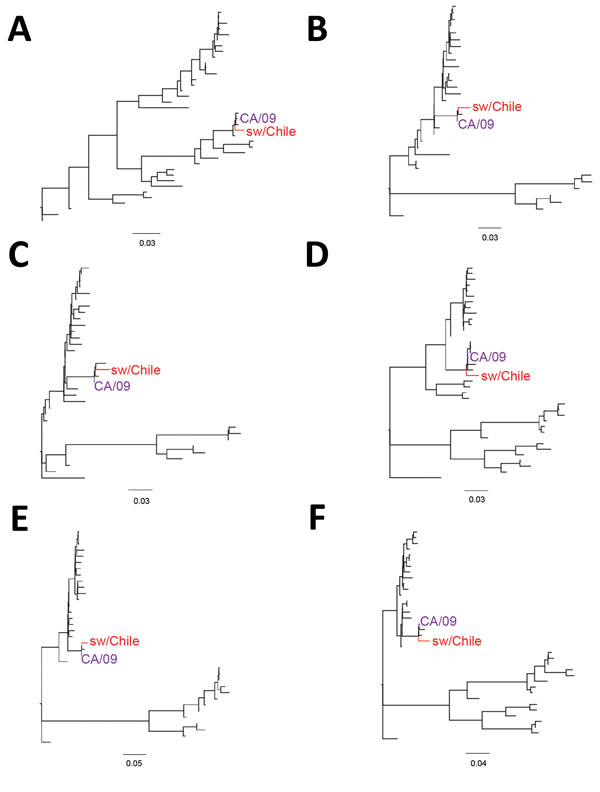
Figure 2. Phylogenetic trees comparing the internal genes of swine influenza virus (H1N2) from Chile (red) and reference viruses. We performed phylogenetic analysis for the matrix (A), nucleoprotein (B), nonstructural (C), polymerase acid (D), polymerase basic (PB) 1 (E), and PB2 (F) gene segments by using RAxML with 200-bootstraps replicates (21). Purple indicates location of control influenza A(H1N1)pdm09 CA/09 virus. Scale bars indicate number of substitutions per site. Detailed phylogenetic trees for these genes are provided in Technical Appendix Figures 1–6.
References
- Komadina N, McVernon J, Hall R, Leder K. A historical perspective of influenza A(H1N2) virus. Emerg Infect Dis. 2014;20:6–12. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Reported infections with variant influenza viruses in the United States since 2005. 2016 Sep 12 [cited 2016 Oct 10]. http://www.cdc.gov/flu/swineflu/variant-cases-us.htm
- Herriman R. Swine flu, H1N2v virus infections reported in Wisconsin, Minnesota. 2016 Jul 1 [cited 2016 Oct 10]. http://outbreaknewstoday.com/swine-flu-h1n2v-virus-infections-reported-in-wisconsin-minnesota-22255/
- Centers for Disease Control and Prevention. Influenza A (H3N2) variant virus [cited 2016 Oct 6]. http://www.cdc.gov/flu/swineflu/h3n2v-cases.htm
- Centers for Disease Control and Prevention. H1N2 variant virus detected in Minnesota. 2012 Sep 7 [cited 2016 Oct 6]. http://www.cdc.gov/flu/spotlights/h1n2v-cases-mn.htm
- Trombetta C, Piccirella S, Perini D, Kistner O, Montomoli E. Emerging influenza strains in the last two decades: a threat of a new pandemic? Vaccines (Basel). 2015;3:172–85. DOIPubMedGoogle Scholar
- Choi MJ, Morin CA, Scheftel J, Vetter SM, Smith K, Lynfield R; Variant Influenza Investigation Team. Variant influenza associated with live animal markets, Minnesota. Zoonoses Public Health. 2015;62:326–30. DOIPubMedGoogle Scholar
- Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, Garten RJ, et al.; Novel Swine-Origin Influenza A (H1N1) Virus Investigation Team. Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med. 2009;360:2605–15. DOIPubMedGoogle Scholar
- Nelson M, Culhane MR, Rovira A, Torremorell M, Guerrero P, Norambuena J. Novel human-like influenza A viruses circulate in swine in Mexico and Chile. PLoS Curr. 2015;7:7.PubMedGoogle Scholar
- Hamilton-West C, Rojas H, Pinto J, Orozco J, Hervé-Claude LP, Urcelay S. Characterization of backyard poultry production systems and disease risk in the central zone of Chile. Res Vet Sci. 2012;93:121–4. DOIPubMedGoogle Scholar
- Bravo-Vasquez N, Di Pillo F, Lazo A, Jiménez-Bluhm P, Schultz-Cherry S, Hamilton-West C. Presence of influenza viruses in backyard poultry and swine in El Yali wetland, Chile. Prev Vet Med. 2016;134:211–5. DOIPubMedGoogle Scholar
- Karlsson EA, Ciuoderis K, Freiden PJ, Seufzer B, Jones JC, Johnson J, et al. Prevalence and characterization of influenza viruses in diverse species in Los Llanos, Colombia. Emerg Microbes Infect. 2013;2:e20. DOIPubMedGoogle Scholar
- Fulcher ML, Randell SH. Human nasal and tracheo-bronchial respiratory epithelial cell culture. In: Randell HS, Fulcher LM, editors. Epithelial cell culture protocols, 2nd ed. Totowa (NJ): Humana Press; 2013. p. 109–21.
- Dash P, Barnett PV, Denyer MS, Jackson T, Stirling CMA, Hawes PC, et al. Foot-and-mouth disease virus replicates only transiently in well-differentiated porcine nasal epithelial cells. J Virol. 2010;84:9149–60. DOIPubMedGoogle Scholar
- Kaplan BS, Russier M, Jeevan T, Marathe B, Govorkova EA, Russell CJ, et al. Novel highly pathogenic avian A(H5N2) and A(H5N8) influenza viruses of clade 2.3.4.4 from North America have limited capacity for replication and transmission in mammals. mSphere. 2016;1:piie00003-16. DOIPubMedGoogle Scholar
- Jones JC, Turpin EA, Bultmann H, Brandt CR, Schultz-Cherry S. Inhibition of influenza virus infection by a novel antiviral peptide that targets viral attachment to cells. J Virol. 2006;80:11960–7. DOIPubMedGoogle Scholar
- Carlson CM, Turpin EA, Moser LA, O’Brien KB, Cline TD, Jones JC, et al. Transforming growth factor-β: activation by neuraminidase and role in highly pathogenic H5N1 influenza pathogenesis. PLoS Pathog. 2010;6:e1001136. DOIPubMedGoogle Scholar
- Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41:95–8.
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7. DOIPubMedGoogle Scholar
- Bao Y, Bolotov P, Dernovoy D, Kiryutin B, Zaslavsky L, Tatusova T, et al. The influenza virus resource at the National Center for Biotechnology Information. J Virol. 2008;82:596–601. DOIPubMedGoogle Scholar
- Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3. DOIPubMedGoogle Scholar
- Sheridan PA, Paich HA, Handy J, Karlsson EA, Hudgens MG, Sammon AB, et al. Obesity is associated with impaired immune response to influenza vaccination in humans. Int J Obes. 2012;36:1072–7. DOIPubMedGoogle Scholar
- Karlsson EA, Ip HS, Hall JS, Yoon SW, Johnson J, Beck MA, et al. Respiratory transmission of an avian H3N8 influenza virus isolated from a harbour seal. Nat Commun. 2014;5:4791. DOIPubMedGoogle Scholar
- Cline TD, Karlsson EA, Freiden P, Seufzer BJ, Rehg JE, Webby RJ, et al. Increased pathogenicity of a reassortant 2009 pandemic H1N1 influenza virus containing an H5N1 hemagglutinin. J Virol. 2011;85:12262–70. DOIPubMedGoogle Scholar
- Reed LJ, Muench H. A simple methods of estimating fifty percent endpoints. Am J Epidemiol. 1938;27:493–7.
- Morton DB. A systematic approach for establishing humane endpoints. ILAR J. 2000;41:80–6. DOIPubMedGoogle Scholar
- Zhou B, Meliopoulos VA, Wang W, Lin X, Stucker KM, Halpin RA, et al. Reversion of cold-adapted live attenuated influenza vaccine into a pathogenic virus. J Virol. 2016;90:8454–63. DOIPubMedGoogle Scholar
- Barman S, Krylov PS, Fabrizio TP, Franks J, Turner JC, Seiler P, et al. Pathogenicity and transmissibility of North American triple reassortant swine influenza A viruses in ferrets. PLoS Pathog. 2012;8:e1002791. DOIPubMedGoogle Scholar
- Nelson MI, Wentworth DE, Culhane MR, Vincent AL, Viboud C, LaPointe MP, et al. Introductions and evolution of human-origin seasonal influenza a viruses in multinational swine populations. J Virol. 2014;88:10110–9. DOIPubMedGoogle Scholar
- Suzuki Y, Ito T, Suzuki T, Holland RE Jr, Chambers TM, Kiso M, et al. Sialic acid species as a determinant of the host range of influenza A viruses. J Virol. 2000;74:11825–31. DOIPubMedGoogle Scholar
- Ito T, Couceiro JNSS, Kelm S, Baum LG, Krauss S, Castrucci MR, et al. Molecular basis for the generation in pigs of influenza A viruses with pandemic potential. J Virol. 1998;72:7367–73.PubMedGoogle Scholar
- Yang H, Chen Y, Qiao C, Xu C, Yan M, Xin X, et al. Two different genotypes of H1N2 swine influenza virus isolated in northern China and their pathogenicity in animals. Vet Microbiol. 2015;175:224–31. DOIPubMedGoogle Scholar
- Lee JH, Pascua PNQ, Decano AG, Kim SM, Park S-J, Kwon H-I, et al. Evaluation of the zoonotic potential of a novel reassortant H1N2 swine influenza virus with gene constellation derived from multiple viral sources. Infect Genet Evol. 2015;34:378–93. DOIPubMedGoogle Scholar
- Fobian K, Fabrizio TP, Yoon S-W, Hansen MS, Webby RJ, Larsen LE. New reassortant and enzootic European swine influenza viruses transmit efficiently through direct contact in the ferret model. J Gen Virol. 2015;96:1603–12. DOIPubMedGoogle Scholar
- Nelson MI, Schaefer R, Gava D, Cantão ME, Ciacci-Zanella JR. Influenza A viruses of human origin in swine, Brazil. Emerg Infect Dis. 2015;21:1339–47. DOIPubMedGoogle Scholar
- Peng X, Wu H, Xu L, Peng X, Cheng L, Jin C, et al. Molecular characterization of a novel reassortant H1N2 influenza virus containing genes from the 2009 pandemic human H1N1 virus in swine from eastern China. Virus Genes. 2016;52:405–10. DOIPubMedGoogle Scholar
- Liu H, Tao J, Zhang P, Yin X, Ha Z, Zhang C. Pathogenic characteristics of a novel triple-reasserted H1N2 swine influenza virus. Biologicals. 2016;44:252–6. DOIPubMedGoogle Scholar
- Pascua PNQ, Song M-S, Lee JH, Baek YH, Kwon HI, Park S-J, et al. Virulence and transmissibility of H1N2 influenza virus in ferrets imply the continuing threat of triple-reassortant swine viruses. Proc Natl Acad Sci U S A. 2012;109:15900–5. DOIPubMedGoogle Scholar
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
Page created: January 17, 2017
Page updated: January 17, 2017
Page reviewed: January 17, 2017
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.