Volume 32, Number 3—March 2026
Research
Strongyloides Genetic Diversity among Humans, Dogs, and Nonhuman Primates, Central African Republic, 2016–2022
Figure 1
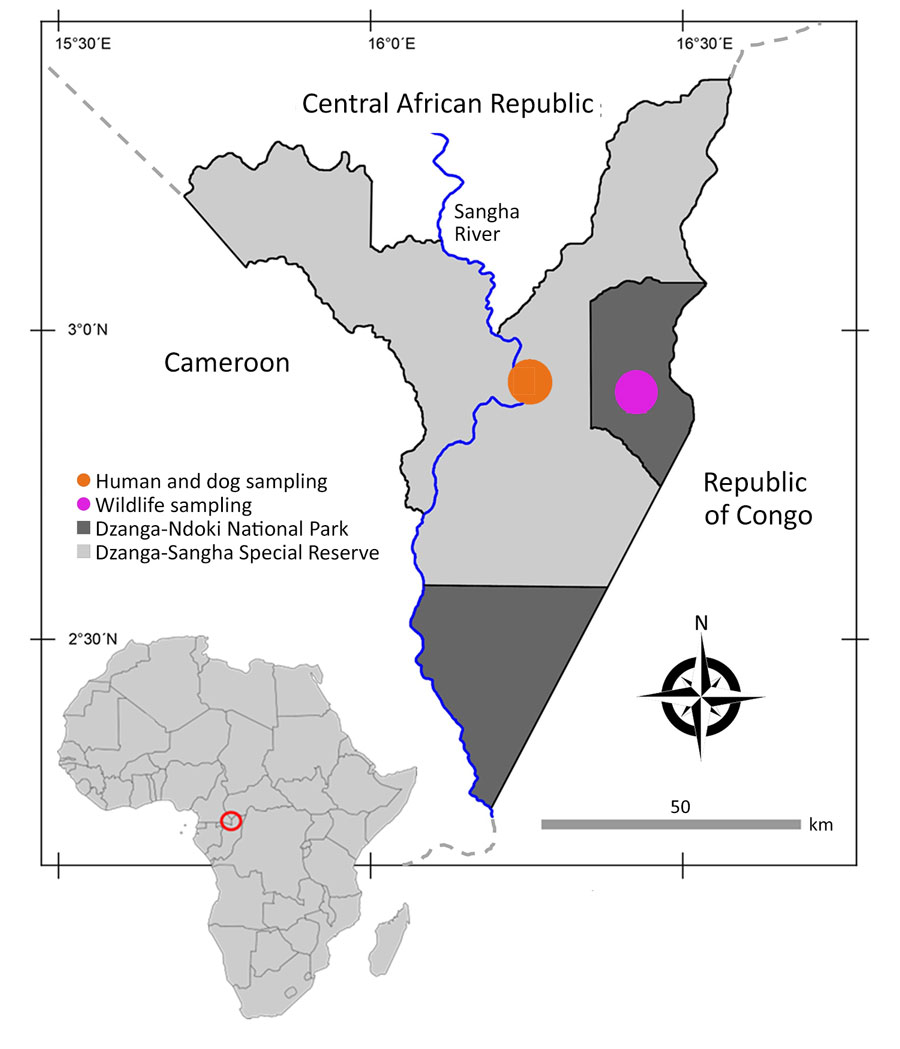
Figure 1. Location of study site in analysis of Strongyloides genetic diversity among humans, dogs, and nonhuman primates, Dzanga-Sangha Protected Areas, Central African Republic, 2016–2022. Inset shows location of Central African Republic in Africa. Figure adapted and adjusted from Hasegawa et al. (18).
References
- Bradbury RS, Pafčo B, Nosková E, Hasegawa H. Strongyloides genotyping: a review of methods and application in public health and population genetics. Int J Parasitol. 2021;51:1153–66. DOIPubMedGoogle Scholar
- Asundi A, Beliavsky A, Liu XJ, Akaberi A, Schwarzer G, Bisoffi Z, et al. Prevalence of strongyloidiasis and schistosomiasis among migrants: a systematic review and meta-analysis. Lancet Glob Health. 2019;7:e236–48. DOIPubMedGoogle Scholar
- Buonfrate D, Bisanzio D, Giorli G, Odermatt P, Fürst T, Greenaway C, et al. The global prevalence of Strongyloides stercoralis infection. Pathogens. 2020;9:468. DOIPubMedGoogle Scholar
- Nutman TB. Human infection with Strongyloides stercoralis and other related Strongyloides species. Parasitology. 2017;144:263–73. DOIPubMedGoogle Scholar
- Jaleta TG, Zhou S, Bemm FM, Schär F, Khieu V, Muth S, et al. Different but overlapping populations of Strongyloides stercoralis in dogs and humans-dogs as a possible source for zoonotic strongyloidiasis. PLoS Negl Trop Dis. 2017;11:
e0005752 . DOIPubMedGoogle Scholar - Nagayasu E, Aung MPPTHH, Hortiwakul T, Hino A, Tanaka T, Higashiarakawa M, et al. A possible origin population of pathogenic intestinal nematodes, Strongyloides stercoralis, unveiled by molecular phylogeny. Sci Rep. 2017;7:4844. DOIPubMedGoogle Scholar
- Gordon CA, Utzinger J, Muhi S, Becker SL, Keiser J, Khieu V, et al. Strongyloidiasis. Nat Rev Dis Primers. 2024;10:6. DOIPubMedGoogle Scholar
- Buonfrate D, Requena-Mendez A, Angheben A, Cinquini M, Cruciani M, Fittipaldo A, et al. Accuracy of molecular biology techniques for the diagnosis of Strongyloides stercoralis infection-A systematic review and meta-analysis. PLoS Negl Trop Dis. 2018;12:
e0006229 . DOIPubMedGoogle Scholar - Kuehne A, Roberts L. Learning from health information challenges in the Central African Republic: where documenting health and humanitarian needs requires fresh approaches. Confl Health. 2021;15:68. DOIPubMedGoogle Scholar
- United Nations Development Programme. Human development insights [cited 2025 Feb 6]. https://hdr.undp.org/data-center/country-insights#/ranks
- Blom A, Yamindou J, Prins HHT. Status of the protected areas of the Central African Republic. Biol Conserv. 2004;118:479–87. DOIGoogle Scholar
- David PM, Nakouné E, Giles-Vernick T. Hotspot or blind spot? Historical perspectives on surveillance and response to epidemics in the Central African Republic. Int J Public Health. 2020;65:241–8. DOIPubMedGoogle Scholar
- Pedersen AB, Davies TJ. Cross-species pathogen transmission and disease emergence in primates. Ecohealth. 2009;6:496–508. DOIPubMedGoogle Scholar
- Hasegawa H, Kalousova B, McLennan MR, Modry D, Profousova-Psenkova I, Shutt-Phillips KA, et al. Strongyloides infections of humans and great apes in Dzanga-Sangha Protected Areas, Central African Republic and in degraded forest fragments in Bulindi, Uganda. Parasitol Int. 2016;65(5 Pt A):367–70. DOIPubMedGoogle Scholar
- Wolfe ND, Dunavan CP, Diamond J. Origins of major human infectious diseases. Nature. 2007;447:279–83. DOIPubMedGoogle Scholar
- World Health Organization. Soil-transmitted helminthiases [cited 2025 Feb 6]. https://www.who.int/data/gho/data/themes/topics/soil-transmitted-helminthiases.
- Pampiglione S, Ricciardi M. Parasitological survey on Pygmies in Central Africa. I. Babinga group (Central African Republic). Riv Parassitol. 1974;35:161–88.
- Hasegawa H, Modrý D, Kitagawa M, Shutt KA, Todd A, Kalousová B, et al. Humans and great apes cohabiting the forest ecosystem in central african republic harbour the same hookworms. PLoS Negl Trop Dis. 2014;8:
e2715 . DOIPubMedGoogle Scholar - Verweij JJ, Canales M, Polman K, Ziem J, Brienen EA, Polderman AM, et al. Molecular diagnosis of Strongyloides stercoralis in faecal samples using real-time PCR. Trans R Soc Trop Med Hyg. 2009;103:342–6. DOIPubMedGoogle Scholar
- Pafčo B, Čížková D, Kreisinger J, Hasegawa H, Vallo P, Shutt K, et al. Metabarcoding analysis of strongylid nematode diversity in two sympatric primate species. Sci Rep. 2018;8:5933. DOIPubMedGoogle Scholar
- Barratt JLN, Lane M, Talundzic E, Richins T, Robertson G, Formenti F, et al. A global genotyping survey of Strongyloides stercoralis and Strongyloides fuelleborni using deep amplicon sequencing. PLoS Negl Trop Dis. 2019;13:
e0007609 . DOIPubMedGoogle Scholar - Jiang H, Lei R, Ding SW, Zhu S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinformatics. 2014;15:182. DOIPubMedGoogle Scholar
- Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3. DOIPubMedGoogle Scholar
- Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7. DOIPubMedGoogle Scholar
- Bradbury RS, Streit A. Is strongyloidiasis a zoonosis from dogs? Philos Trans R Soc Lond B Biol Sci. 2024;379:
20220445 . DOIPubMedGoogle Scholar - Lapat JJ, Opee J, Apio MC, Akello S, Ojul CL, Onekalit R, et al. A One Health approach toward the control and elimination of soil-transmitted helminthic infections in endemic areas. IJID One Health. 2024;2:100021. DOIGoogle Scholar
- Nosková E, Sambucci KM, Petrželková KJ, Červená B, Modrý D, Pafčo B. Strongyloides in non-human primates: significance for public health control. Philos Trans R Soc Lond B Biol Sci. 2024;379:
20230006 . DOIPubMedGoogle Scholar - Beknazarova M, Barratt JLN, Bradbury RS, Lane M, Whiley H, Ross K. Detection of classic and cryptic Strongyloides genotypes by deep amplicon sequencing: A preliminary survey of dog and human specimens collected from remote Australian communities. PLoS Negl Trop Dis. 2019;13:
e0007241 . DOIPubMedGoogle Scholar - Nijsse R, Mughini-Gras L, Wagenaar JA, Ploeger HW. Coprophagy in dogs interferes in the diagnosis of parasitic infections by faecal examination. Vet Parasitol. 2014;204:304–9. DOIPubMedGoogle Scholar
- Sandground JH. Biological studies on the life-cycle in the genus Strongyloides grassi, 1879. Am J Epidemiol. 1926;6:337–88. DOIGoogle Scholar
- Richins T, Sapp SGH, Ketzis JK, Willingham AL, Mukaratirwa S, Qvarnstrom Y, et al. Genetic characterization of Strongyloides fuelleborni infecting free-roaming African vervets (Chlorocebus aethiops sabaeus) on the Caribbean island of St. Kitts. Int J Parasitol Parasites Wildl. 2023;20:153–61. DOIPubMedGoogle Scholar
- Alum A, Rubino JR, Ijaz MK. The global war against intestinal parasites—should we use a holistic approach? Int J Infect Dis. 2010;14:e732–8. DOIPubMedGoogle Scholar
- Horton J. Global anthelmintic chemotherapy programs: learning from history. Trends Parasitol. 2003;19:405–9. DOIPubMedGoogle Scholar
- A F White M, Whiley H, E Ross K. A review of Strongyloides spp. environmental sources worldwide. Pathogens. 2019;8:8.PubMedGoogle Scholar
- Khieu V, Hattendorf J, Schär F, Marti H, Char MC, Muth S, et al. Strongyloides stercoralis infection and re-infection in a cohort of children in Cambodia. Parasitol Int. 2014;63:708–12. DOIPubMedGoogle Scholar
- Kamgno J, Boussinesq M, Labrousse F, Nkegoum B, Thylefors BI, Mackenzie CD. Encephalopathy after ivermectin treatment in a patient infected with Loa loa and Plasmodium spp. Am J Trop Med Hyg. 2008;78:546–51. DOIPubMedGoogle Scholar
- Zinsstag J, Kaiser-Grolimund A, Heitz-Tokpa K, Sreedharan R, Lubroth J, Caya F, et al. Advancing One human-animal-environment Health for global health security: what does the evidence say? Lancet. 2023;401:591–604. DOIPubMedGoogle Scholar
Page created: February 07, 2026
Page updated: May 04, 2026
Page reviewed: May 04, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.