Volume 32, Number 3—March 2026
Research
Strongyloides Genetic Diversity among Humans, Dogs, and Nonhuman Primates, Central African Republic, 2016–2022
Figure 2
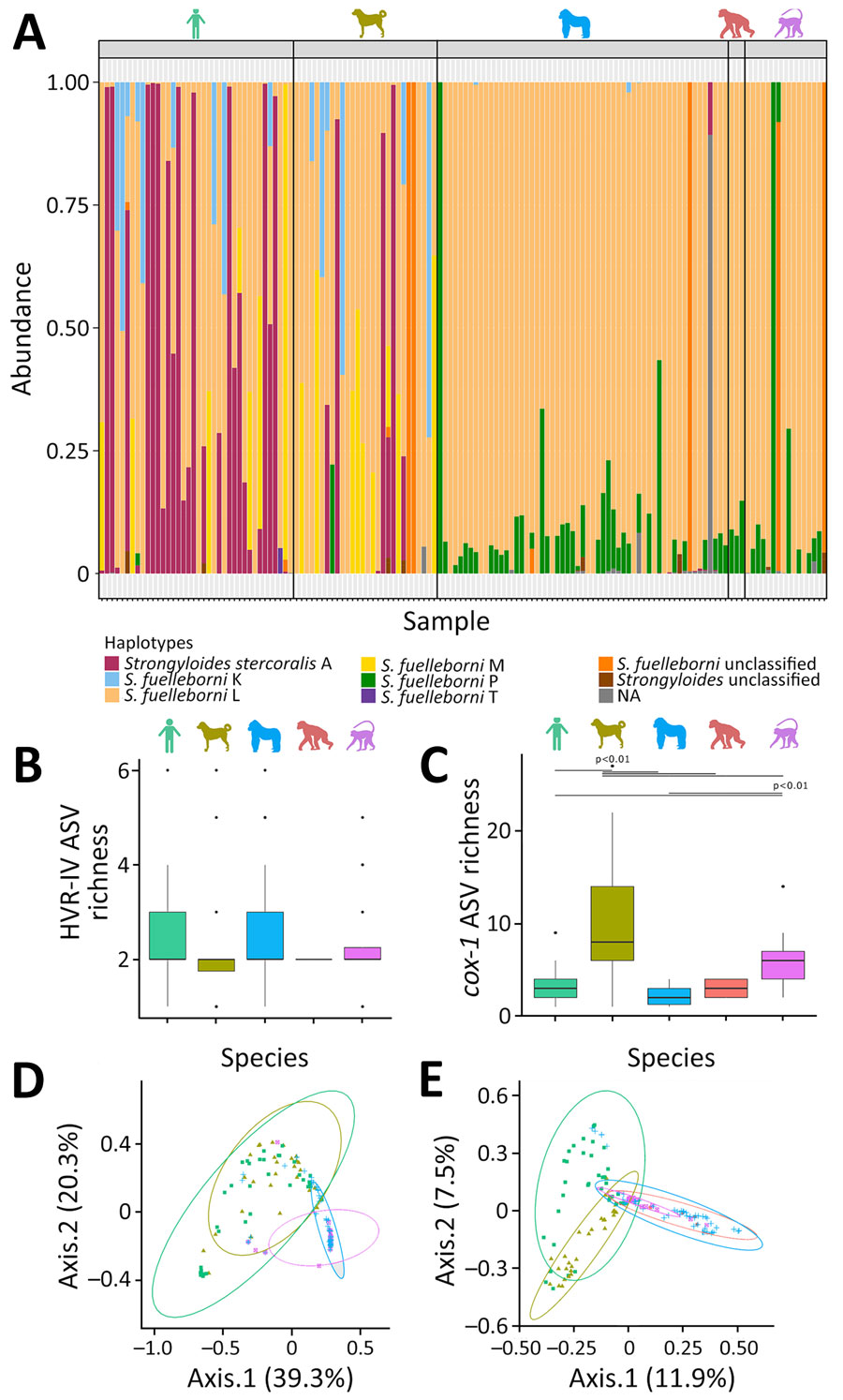
Figure 2. Relative community composition of Strongyloides HVR-IV-18S rRNA haplotypes across examined hosts in study of Strongyloides genetic diversity among humans, dogs, and nonhuman primates, Dzanga-Sangha Protected Areas, Central African Republic, 2016–2022. A) Relative abundance of haplotypes shown as color panels; each column represents a single sample. B) Boxplot showing the α diversity of Strongyloides HVR-IV 18S rRNA haplotypes, represented by the number of ASVs per sample (dots) grouped by host species. C) Boxplot showing the α diversity of Strongyloides cox1 haplotypes represented by the number of ASVs per sample (dots) grouped by host species. D) Beta diversity of Strongyloides HVR-IV 18S rRNA haplotype communities, based on Bray-Curtis ecologic distances (relative abundance of reads), visualized using principal coordinate analysis ordination diagrams. E) Principal coordinate analysis showing the β diversity of Strongyloides cox1 haplotypes. Color silhouettes (A-C) and data points (D, E) indicate host species: light green, human; olive green, dog; blue, gorilla; red, chimpanzee; violet, mangabey. ASV, amplicon sequencing variant; HVR-IV, hypervariable region IV.