Strongyloides Genetic Diversity among Humans, Dogs, and Nonhuman Primates, Central African Republic, 2016–2022
Eva Nosková, Vladislav Ilík, Frédéric Stéphane Singa Niatou, Laurent Dumas, Terence Fuh, Jean-Francais Dicky, Terézia Kurucová, Vojtech Baláž, Klára Judita Petrželková, and Barbora Pafčo

Author affiliation: Department of Botany and Zoology, Masaryk University, Brno, Czech Republic (E. Nosková, V. Ilík, B. Pafčo); Masaryk University, Central European Institute of Technology, Brno (T. Kurucová); Czech Academy of Sciences, Institute of Vertebrate Biology, Brno (E. Nosková, V. Ilík, K.J. Petrželková, B. Pafčo); Czech Academy of Sciences, Biology Center, Institute of Parasitology, Ceske Budejovice, Czech Republic (K.J. Petrželková); WWF Central African Republic Programme Office, Dzanga Sangha Protected Areas, Bangui, Central African Republic (F.S. Singa Niatou, T. Fuh); Toulouse National School of Veterinarians, Toulouse, France (L. Dumas); University of Veterinary Sciences, Brno (V. Baláž)
Main Article
Figure 3
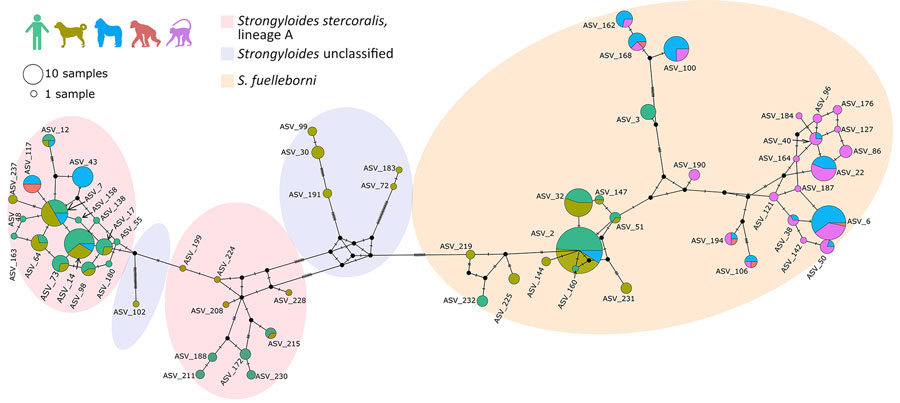
Figure 3. Median-joining Strongyloides haplotype network for the mitochondrial cox1 gene studied in various host species in study of Strongyloides genetic diversity among humans, dogs, and nonhuman primates, Dzanga-Sangha Protected Areas, Central African Republic, 2016–2022. Each circle represents 1 haplotype; circle size indicates the number of hosts harboring the respective haplotype. The colors inside the circles indicate the host species: light green, human; olive green, dog; blue, gorilla; red, chimpanzee; violet, mangabey. The hatch marks beside the branches indicate the number of mutation steps between the haplotypes. Missing haplotypes are indicated by small black circles. Shaded ovals indicate species/lineage type. ASV, amplicon sequencing variant.
Main Article
Page created: February 07, 2026
Page updated: May 04, 2026
Page reviewed: May 04, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
