Volume 24, Number 1—January 2018
Research
Characterization of a Feline Influenza A(H7N2) Virus
Figure 1
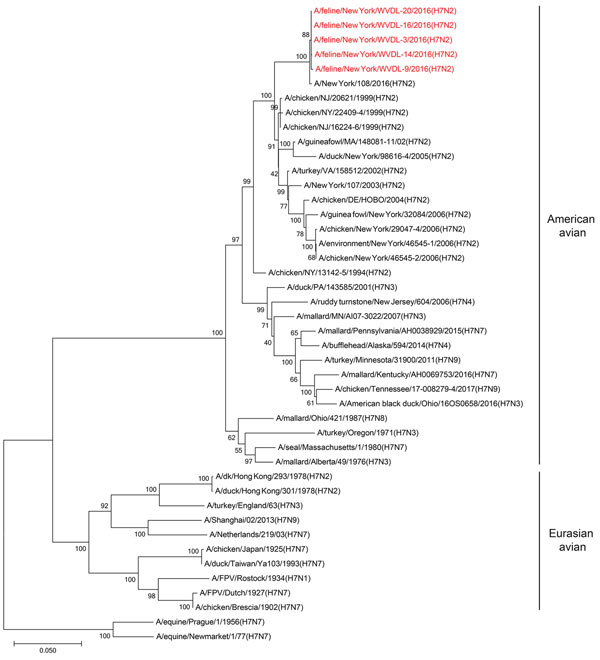
Figure 1. Phylogenetic tree of influenza A viral HA segments. Phylogenetic analysis was performed for selected influenza A viruses representing major lineages. The evolutionary history was inferred using the neighbor-joining method (12). The optimal tree with the branch length sum of 1.22521320 is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) is shown next to the branches (13). The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Tamura 3-parameter method (14) and are in the units of the number of base substitutions per site. The analysis involved 44 nt sequences. Codon positions included were 1st + 2nd + 3rd + noncoding. All positions containing gaps and missing data were eliminated. The final dataset contained a total of 1,612 positions. Evolutionary analyses were conducted in MEGA7 (15).
References
- Wright PF, Neumann G, Kawaoka Y. Orthomyxoviruses. In: Knipe DM, Howley PM, Cohen JI, Griffin DE, Lamb RA, Martin MA, et al., editors. Fields Virology. Sixth ed. Philadelphia, PA, USA: Lippincott Williams & Wilkins; 2013. p. 1186–243.
- Crawford PC, Dubovi EJ, Castleman WL, Stephenson I, Gibbs EP, Chen L, et al. Transmission of equine influenza virus to dogs. Science. 2005;310:482–5. DOIPubMedGoogle Scholar
- Xie X, Ma K, Liu Y. Influenza A virus infection in dogs: epizootiology, evolution and prevention - A review. Acta Vet Hung. 2016;64:125–39. DOIPubMedGoogle Scholar
- Song D, Kang B, Lee C, Jung K, Ha G, Kang D, et al. Transmission of avian influenza virus (H3N2) to dogs. Emerg Infect Dis. 2008;14:741–6. DOIPubMedGoogle Scholar
- Lee YN, Lee DH, Lee HJ, Park JK, Yuk SS, Sung HJ, et al. Evidence of H3N2 canine influenza virus infection before 2007. Vet Rec. 2012;171:477. DOIPubMedGoogle Scholar
- Fiorentini L, Taddei R, Moreno A, Gelmetti D, Barbieri I, De Marco MA, et al. Influenza A pandemic (H1N1) 2009 virus outbreak in a cat colony in Italy. Zoonoses Public Health. 2011;58:573–81. DOIPubMedGoogle Scholar
- Belser JA, Pulit-Penaloza JA, Sun X, Brock N, Pappas C, Creager HM, et al. A novel A(H7N2) influenza virus isolated from a veterinarian caring for cats in a New York City animal shelter causes mild disease and transmits poorly in the ferret model. J Virol. 2017;91:e00672–17. DOIPubMedGoogle Scholar
- Spackman E, Senne DA, Davison S, Suarez DL. Sequence analysis of recent H7 avian influenza viruses associated with three different outbreaks in commercial poultry in the United States. J Virol. 2003;77:13399–402. DOIPubMedGoogle Scholar
- Imai M, Watanabe T, Hatta M, Das SC, Ozawa M, Shinya K, et al. Experimental adaptation of an influenza H5 HA confers respiratory droplet transmission to a reassortant H5 HA/H1N1 virus in ferrets. Nature. 2012;486:420–8.PubMedGoogle Scholar
- Watanabe T, Kiso M, Fukuyama S, Nakajima N, Imai M, Yamada S, et al. Characterization of H7N9 influenza A viruses isolated from humans. Nature. 2013;501:551–5. DOIPubMedGoogle Scholar
- Arafa AS, Yamada S, Imai M, Watanabe T, Yamayoshi S, Iwatsuki-Horimoto K, et al. Risk assessment of recent Egyptian H5N1 influenza viruses. Sci Rep. 2016;6:38388. DOIPubMedGoogle Scholar
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.PubMedGoogle Scholar
- Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–91. DOIPubMedGoogle Scholar
- Tamura K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol Biol Evol. 1992;9:678–87.PubMedGoogle Scholar
- Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. H5N1 genetic changes inventory:
- a tool for influenza surveillance and preparedness. 2012 [cited 2017 Jul 27]. https://www.cdc.gov/flu/pdf/avianflu/h5n1-inventory.pdf
- Connor RJ, Kawaoka Y, Webster RG, Paulson JC. Receptor specificity in human, avian, and equine H2 and H3 influenza virus isolates. Virology. 1994;205:17–23. DOIPubMedGoogle Scholar
- Gambaryan AS, Tuzikov AB, Piskarev VE, Yamnikova SS, Lvov DK, Robertson JS, et al. Specification of receptor-binding phenotypes of influenza virus isolates from different hosts using synthetic sialylglycopolymers: non-egg-adapted human H1 and H3 influenza A and influenza B viruses share a common high binding affinity for 6′-sialyl(N-acetyllactosamine). Virology. 1997;232:345–50. DOIPubMedGoogle Scholar
- Matrosovich MN, Gambaryan AS, Teneberg S, Piskarev VE, Yamnikova SS, Lvov DK, et al. Avian influenza A viruses differ from human viruses by recognition of sialyloligosaccharides and gangliosides and by a higher conservation of the HA receptor-binding site. Virology. 1997;233:224–34. DOIPubMedGoogle Scholar
- Wang H, Wu X, Cheng Y, An Y, Ning Z. Tissue distribution of human and avian type sialic acid influenza virus receptors in domestic cat. Acta Vet Hung. 2013;61:537–46. DOIPubMedGoogle Scholar
- Said AW, Usui T, Shinya K, Ono E, Ito T, Hikasa Y, et al. A Sero-survey of subtype H3 influenza a virus infection in dogs and cats in Japan. J Vet Med Sci. 2011;73:541–4. DOIPubMedGoogle Scholar
- Thongratsakul S, Suzuki Y, Hiramatsu H, Sakpuaram T, Sirinarumitr T, Poolkhet C, et al. Avian and human influenza A virus receptors in trachea and lung of animals. Asian Pac J Allergy Immunol. 2010;28:294–301.PubMedGoogle Scholar
- Harder TC, Vahlenkamp TW. Influenza virus infections in dogs and cats. Vet Immunol Immunopathol. 2010;134:54–60. DOIPubMedGoogle Scholar
- Fouchier RA, Schneeberger PM, Rozendaal FW, Broekman JM, Kemink SA, Munster V, et al. Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome. Proc Natl Acad Sci U S A. 2004;101:1356–61. DOIPubMedGoogle Scholar
- Koopmans M, Wilbrink B, Conyn M, Natrop G, van der Nat H, Vennema H, et al. Transmission of H7N7 avian influenza A virus to human beings during a large outbreak in commercial poultry farms in the Netherlands. Lancet. 2004;363:587–93. DOIPubMedGoogle Scholar
1These authors contributed equally to this article.