Volume 27, Number 3—March 2021
Research
Daily Forecasting of Regional Epidemics of Coronavirus Disease with Bayesian Uncertainty Quantification, United States
Figure 4
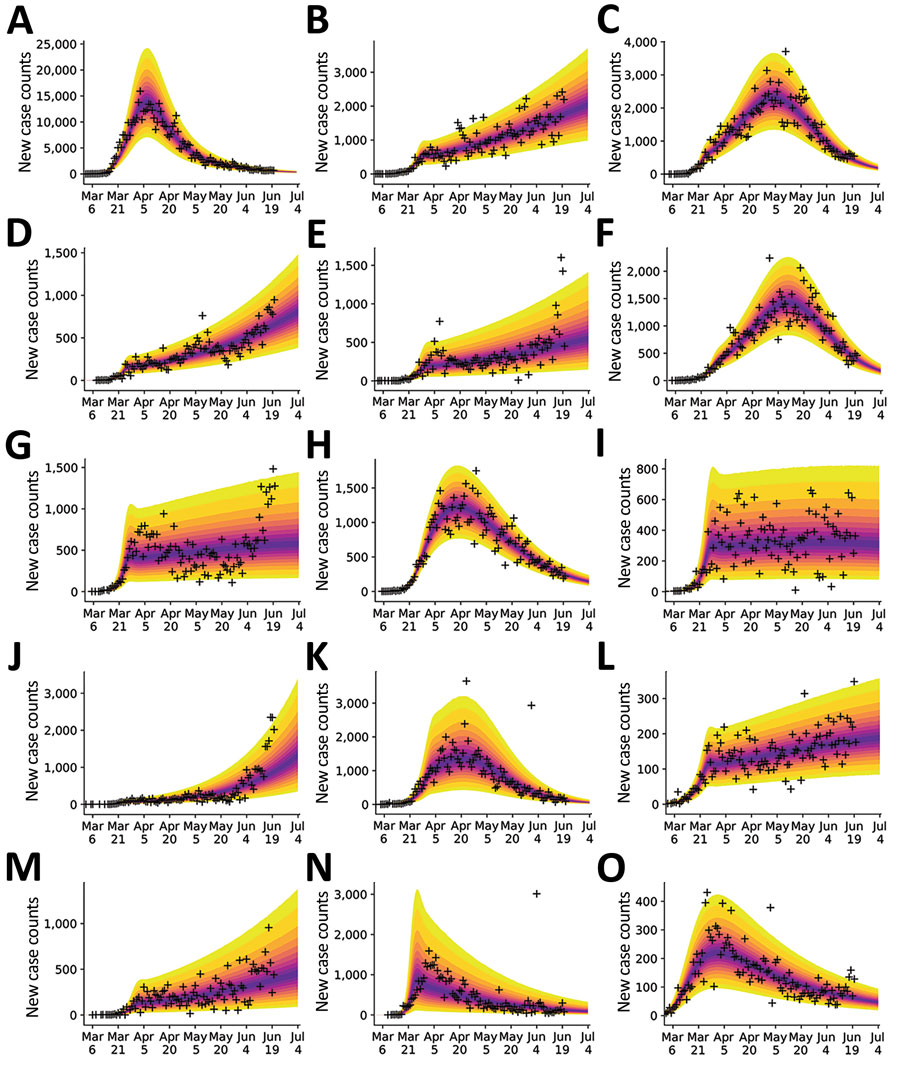
Figure 4. Bayesian predictive inferences for daily new case counts of coronavirus disease in the 15 most populous metropolitan statistical areas, United States, March 1–June 21, 2020. Predictions conditioned on the compartmental model with structure defined by n = 0, which accounts for a single initial period of social distancing. Inferences shown for the metropolitan statistical areas for the following cities: A) New York City, New York; B) Los Angeles, California; C) Chicago, Illinois; D) Dallas, Texas; E) Houston, Texas; F) Washington, DC; G) Miami, Florida; H) Philadelphia, Pennsylvania; I) Atlanta, Georgia; J) Phoenix, Arizona; K) Boston, Massachusetts; L) San Francisco, California; M) Riverside, California; N) Detroit, Michigan; and O) Seattle, Washington. Crosses indicate observed daily case reports. The shaded region indicates the 95% credible interval for predictions of daily case reports. The color-coded bands within the shaded region indicate alternate credible intervals. The model was parametrized with uncertainty quantification by using data from January 21–June 21, 2020. The uncertainty bands/inferred model was used to make predictions for 14 days after the last observed data: the last prediction date was July 5, 2020.